| DPOGS202404 | ||
|---|---|---|
| Transcript | DPOGS202404-TA | 582 bp |
| Protein | DPOGS202404-PA | 133 aa |
| Genomic position | DPSCF300233 - 172492-180950 | |
| RNAseq coverage | 6x (Rank: top 87%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | % | |||
| Bombyx | % | |||
| Drosophila | CG34384-PE | 1e-10 | 56.14% | |
| EBI UniRef50 | UniRef50_Q86XP1 | 4e-15 | 57.33% | Diacylglycerol kinase eta n=149 Tax=Chordata RepID=DGKH_HUMAN |
| NCBI RefSeq | XP_002734721.1 | 1e-17 | 54.35% | PREDICTED: diacylglycerol kinase, eta-like [Saccoglossus kowalevskii] |
| NCBI nr blastp | gi|291229516 | 3e-16 | 54.35% | PREDICTED: diacylglycerol kinase, eta-like [Saccoglossus kowalevskii] |
| NCBI nr blastx | gi|291229516 | 2e-17 | 50.44% | PREDICTED: diacylglycerol kinase, eta-like [Saccoglossus kowalevskii] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 1.3e-10 | protein binding | |
| KEGG pathway | rno:361076 | 2e-15 | ||
| K00901 (E2.7.1.107, DGK, dgkA) | maps-> | Phosphatidylinositol signaling system | ||
| Glycerolipid metabolism | ||||
| Glycerophospholipid metabolism | ||||
| InterPro domain | [89-128] IPR011993 | 1.3e-10 | Pleckstrin homology-type | |
| [93-128] IPR001849 | 2.1e-07 | Pleckstrin homology domain | ||
| Orthology group | ||||
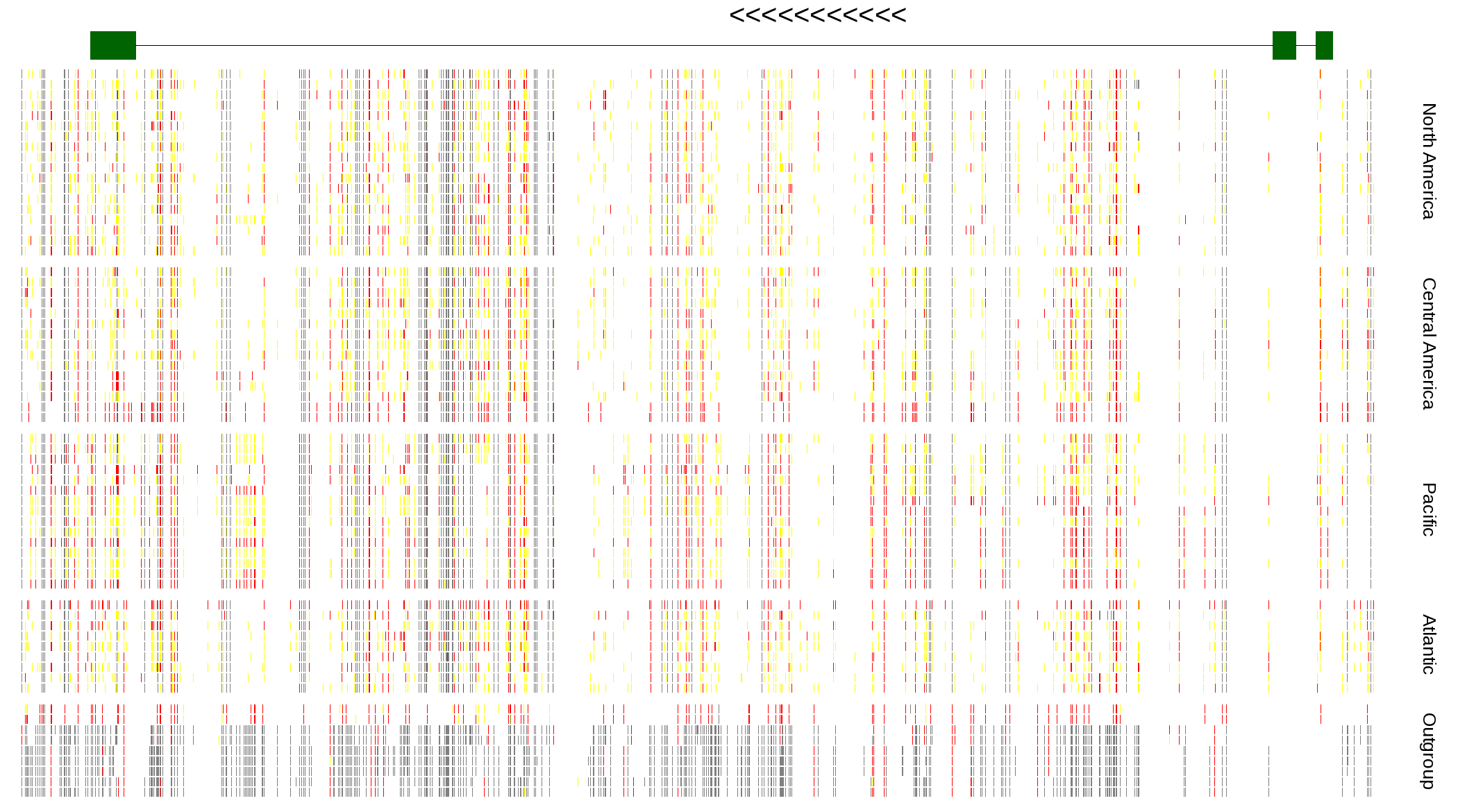
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202404-TA
ATGGCGCCACCTAGAGACCATTTTCACATACCCGTCCATTGGAAAAACTTTCAGGAACATTCTTGTTGCTTGTTCAAATTTGTACTGATAGATGTCGCTAGCTCTGGTCGGCTTGGAGCGAGCGCCGGCGGCGCGATGTCAATAGCTGAAGGTAATGCGACCGGCCACGAAAGGAAAACAGCACCCGGAGAGGATAGCACGGATAGTGACAATGAAGCTGAGAGAAAAGCTCTTCTACGCAGGATATCCACCAGCAAATGTGTCGGCAGACTGCCGGTCGTGAAGGAGGGTTACTTGATGAAGCAGACACGAAGCTTCCAGCGATGGCAGCGACGTTACTTCAAACTGCGAGGTCGGACCCTCTACTACGCCAAGAAGAAAGACGTAAGTGATGAGATCTAAGACCTTCGCGAGTATTCATTTAAACGAAACTAATATTGTCGGATTACTACGCGAATTTTATTATTTTTAAAACTACATACTCCCGACGTTTCGGTTACTTCGCAGCAACCGTGATCACGGGCGGACGAGATGATATTCTCAATAACGTGAAATAATCCGAAAATATTAGTTTCCTTTTAA
>DPOGS202404-PA
MAPPRDHFHIPVHWKNFQEHSCCLFKFVLIDVASSGRLGASAGGAMSIAEGNATGHERKTAPGEDSTDSDNEAERKALLRRISTSKCVGRLPVVKEGYLMKQTRSFQRWQRRYFKLRGRTLYYAKKKDVSDEI-