| DPOGS202741 | ||
|---|---|---|
| Transcript | DPOGS202741-TA | 180 bp |
| Protein | DPOGS202741-PA | 59 aa |
| Genomic position | DPSCF300284 + 203745-203924 | |
| RNAseq coverage | 121x (Rank: top 57%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012684 | 8e-22 | 72.88% | |
| Bombyx | BGIBMGA005429-TA | 3e-21 | 74.58% | |
| Drosophila | CG34161-PC | 3e-10 | 50.85% | |
| EBI UniRef50 | UniRef50_D8LWV6 | 3e-11 | 55.93% | Singapore isolate B (sub-type 7) whole genome shotgun sequence assembly, scaffold_0 n=1 Tax=Blastocystis hominis RepID=D8LWV6_BLAHO |
| NCBI RefSeq | XP_625017.1 | 4e-12 | 54.24% | PREDICTED: similar to Acylphosphatase, organ-common type isozyme (Acylphosphate phosphohydrolase) (Acylphosphatase, erythrocyte isozyme) [Apis mellifera] |
| NCBI nr blastp | gi|354499972 | 9e-12 | 54.39% | PREDICTED: acylphosphatase-2-like [Cricetulus griseus] |
| NCBI nr blastx | gi|307191446 | 9e-12 | 55.17% | Acylphosphatase-1 [Camponotus floridanus] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ame:552639 | 1e-11 | ||
| K01512 (E3.6.1.7, acyP) | maps-> | Benzoate degradation via CoA ligation | ||
| Pyruvate metabolism | ||||
| InterPro domain | [1-59] IPR001792 | 1.9e-09 | Acylphosphatase-like | |
| Orthology group | MCL11605 | Multiple-copy universal gene | ||
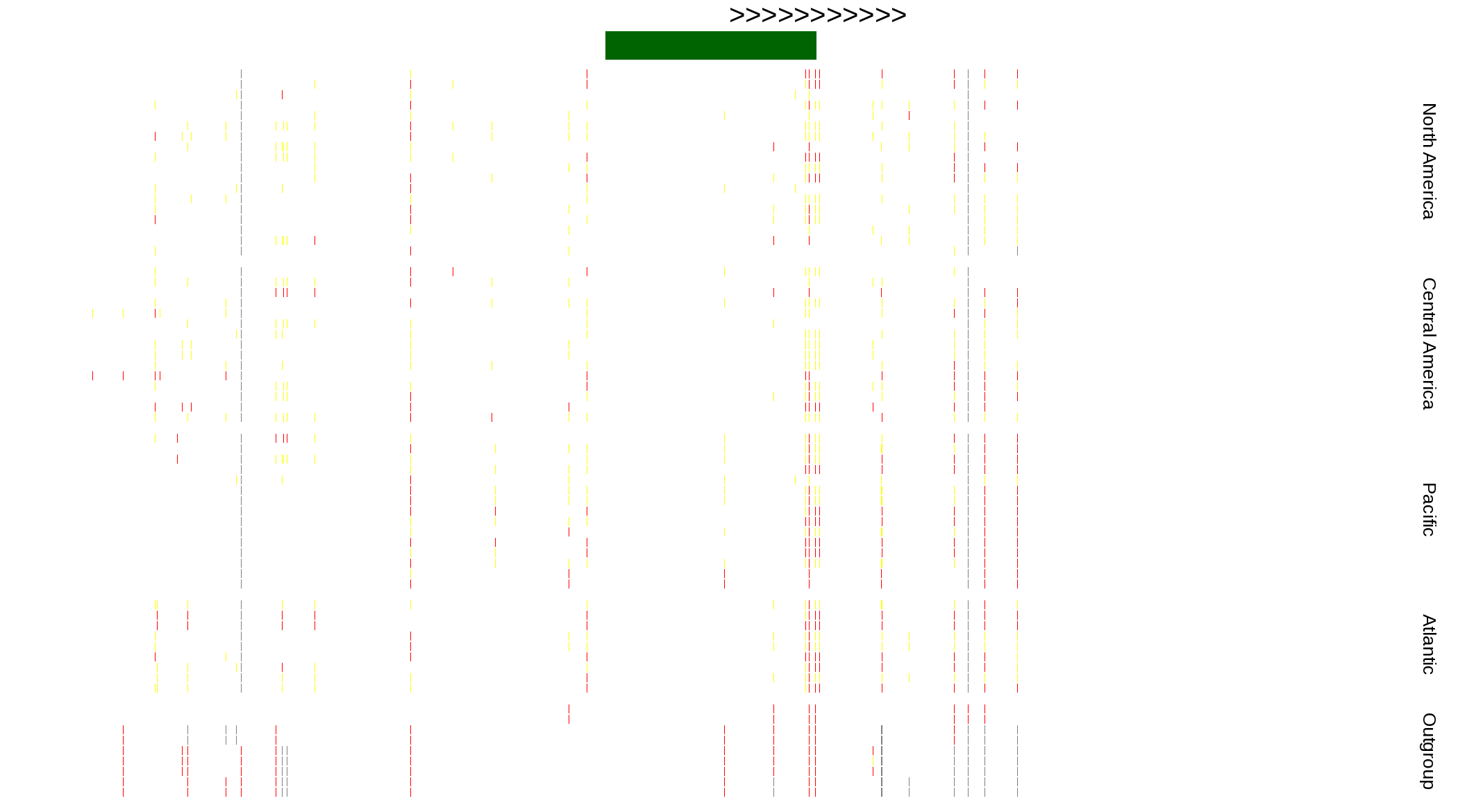
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202741-TA
ATGAATACATCTCAGGGCACCGTTATCGGCCAGATGCAAGGATCCCAAATAGCAATTGACGGAATGAAAGTATGGCTCCAGAAAACCGGCAGCCCCAAATCGAAAATCGAGAAAGCGGATTTTAGGAATGAAGGCCCAATTAATAACTGTGCATTCCGAAATTTCGAAATACGGCGATGA
>DPOGS202741-PA
MNTSQGTVIGQMQGSQIAIDGMKVWLQKTGSPKSKIEKADFRNEGPINNCAFRNFEIRR-