| DPOGS203218 | ||
|---|---|---|
| Transcript | DPOGS203218-TA | 264 bp |
| Protein | DPOGS203218-PA | 87 aa |
| Genomic position | DPSCF300035 + 939686-940023 | |
| RNAseq coverage | 533x (Rank: top 24%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006494 | 2e-43 | 97.70% | |
| Bombyx | BGIBMGA011505-TA | 2e-28 | 93.75% | |
| Drosophila | CG4546-PA | 9e-15 | 46.99% | |
| EBI UniRef50 | UniRef50_Q5DAS4 | 3e-19 | 56.32% | SJCHGC03493 protein n=3 Tax=Eumetazoa RepID=Q5DAS4_SCHJA |
| NCBI RefSeq | XP_624300.1 | 7e-21 | 64.20% | PREDICTED: similar to C-Myc-binding protein (Associate of Myc 1) (AMY-1) [Apis mellifera] |
| NCBI nr blastp | gi|383856719 | 1e-20 | 62.11% | PREDICTED: c-Myc-binding protein-like [Megachile rotundata] |
| NCBI nr blastx | gi|383856719 | 1e-20 | 61.54% | PREDICTED: c-Myc-binding protein-like [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | dme:Dmel_CG4546 | 9e-13 | ||
| K00934 (E2.7.3.3) | maps-> | Arginine and proline metabolism | ||
| Orthology group | MCL16915 | Patchy | ||
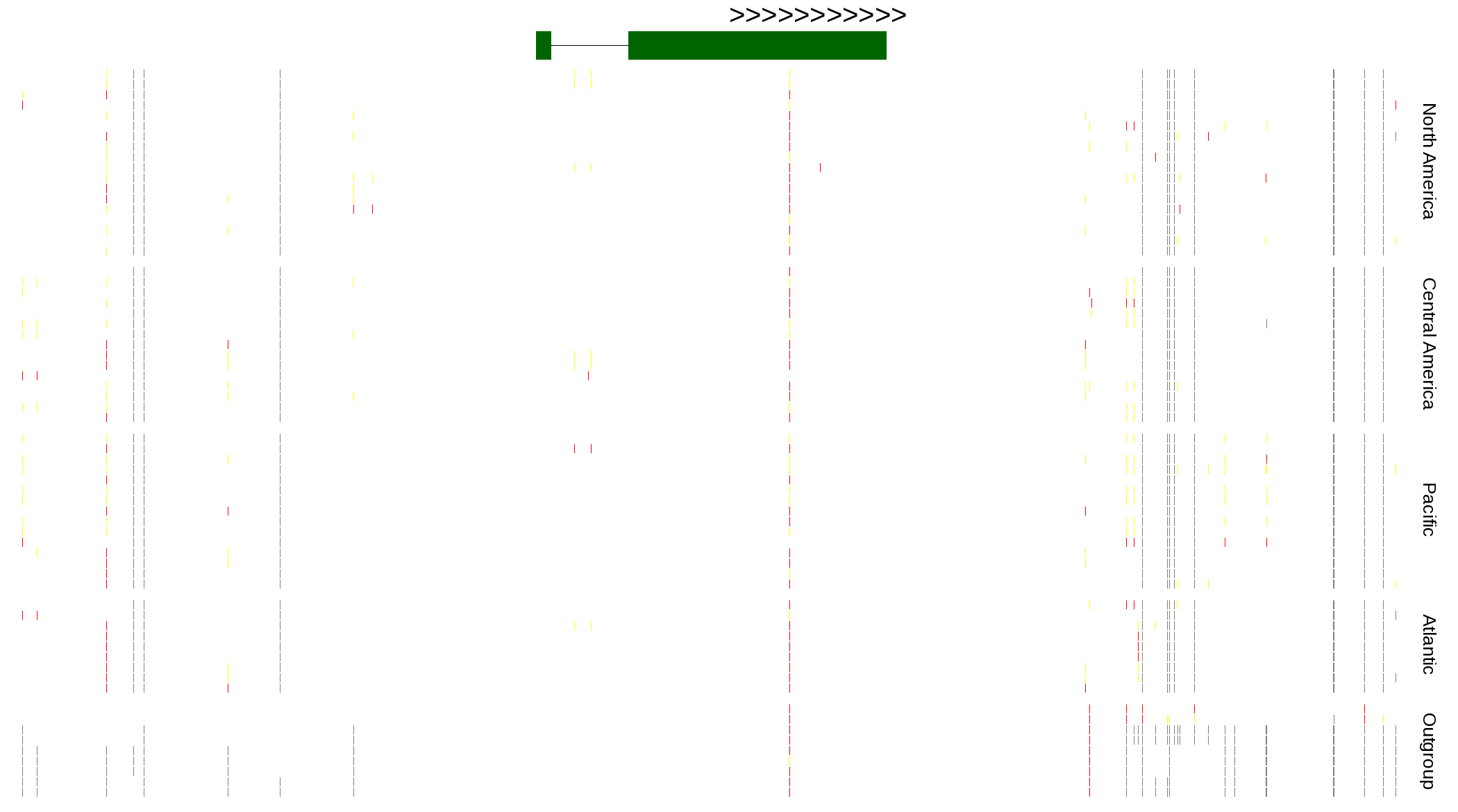
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203218-TA
ATGTCTTCATACAAACCTATTGACTCTAAAAGGGAGGAATTTAGACGTTACCTAGAGCGGGCTGGAGTAATGGACGCCCTTACGAAGGTCCTTGTAAGTCTTTATGAAGAACCGGACAAACCAGAAGATGCCTTAGAATATGTTCGTAAGCATTTAGGAACTGACGGCGGAGAGGATGAGCTAGAAGCAGCACGAGCACGTATAGCAGAATTGGAAGCAGAAAATGCACTCTTAAAAGGAGAAACTGCTCCTACTGAAGGATAA
>DPOGS203218-PA
MSSYKPIDSKREEFRRYLERAGVMDALTKVLVSLYEEPDKPEDALEYVRKHLGTDGGEDELEAARARIAELEAENALLKGETAPTEG-