| DPOGS203325 | ||
|---|---|---|
| Transcript | DPOGS203325-TA | 354 bp |
| Protein | DPOGS203325-PA | 117 aa |
| Genomic position | DPSCF300003 - 428923-429695 | |
| RNAseq coverage | 150x (Rank: top 53%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006200 | 1e-33 | 53.85% | |
| Bombyx | BGIBMGA002087-TA | 2e-34 | 57.39% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_A9QXF1 | 2e-10 | 28.95% | MIF n=2 Tax=root RepID=A9QXF1_HALDV |
| NCBI RefSeq | NP_001040199.1 | 7e-13 | 37.61% | macrophage migration inhibitory factor [Bombyx mori] |
| NCBI nr blastp | gi|114051978 | 2e-11 | 37.61% | macrophage migration inhibitory factor [Bombyx mori] |
| NCBI nr blastx | gi|114051978 | 4e-11 | 37.61% | macrophage migration inhibitory factor [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ecb:100053618 | 2e-09 | ||
| K07253 (MIF) | maps-> | Tyrosine metabolism | ||
| Phenylalanine metabolism | ||||
| InterPro domain | [2-115] IPR014347 | 1.1e-21 | Tautomerase | |
| [2-115] IPR001398 | 1.6e-17 | Macrophage migration inhibitory factor | ||
| Orthology group | MCL26537 | Lepidoptera specific | ||
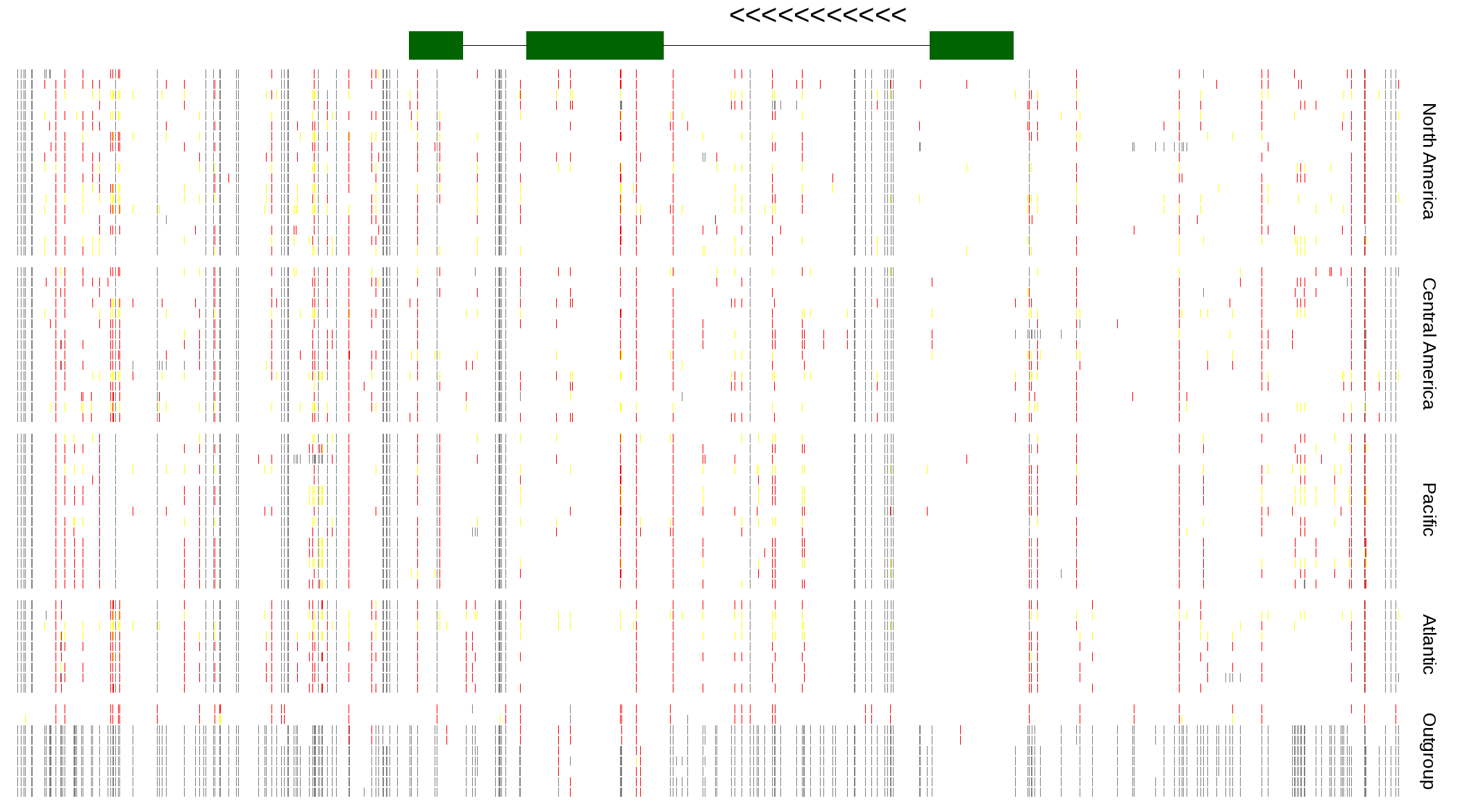
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203325-TA
ATGCCTTGCTTAAATATTTTCACGAACATCGCCAAGTCCCAAATACCACAAAACCTTGTCTCAAAAATTATCCCGGTTCTCGTGAAGGGTGTGAAAAAACCACCTGAGAAATTCATCTGCGTGGTCAATGCCGATTGCTTCTTAAATATGAAGGAAGACTCGGAAACGCCTGGTATCGTAGCCACGCTGGAATCCATAGGACATTTAGGTCCAGAGGAAAATGACATCATTGTTAAGGAGCTTGCGAAATTCTTTGAAACAGAACTTGGCGTGCAACCCAGCAGGTTCTTCCTAACACTTTACGATCTGGAAAAGCACAATGTCGCAATAAACGCAAGCACCATTAAAGCGTAA
>DPOGS203325-PA
MPCLNIFTNIAKSQIPQNLVSKIIPVLVKGVKKPPEKFICVVNADCFLNMKEDSETPGIVATLESIGHLGPEENDIIVKELAKFFETELGVQPSRFFLTLYDLEKHNVAINASTIKA-