| DPOGS204154 | ||
|---|---|---|
| Transcript | DPOGS204154-TA | 600 bp |
| Protein | DPOGS204154-PA | 199 aa |
| Genomic position | DPSCF300034 - 614615-617042 | |
| RNAseq coverage | 510x (Rank: top 25%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004126 | 7e-64 | 91.94% | |
| Bombyx | BGIBMGA005036-TA | 8e-92 | 80.00% | |
| Drosophila | CG15771-PA | 5e-50 | 48.47% | |
| EBI UniRef50 | UniRef50_E2AZQ6 | 5e-52 | 48.53% | Neurochondrin-like protein n=13 Tax=Endopterygota RepID=E2AZQ6_CAMFO |
| NCBI RefSeq | XP_562137.3 | 9e-51 | 47.32% | AGAP004391-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|345495288 | 8e-53 | 50.78% | PREDICTED: neurochondrin homolog [Nasonia vitripennis] |
| NCBI nr blastx | gi|345495288 | 2e-51 | 50.78% | PREDICTED: neurochondrin homolog [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0008152 | 1.9e-15 | metabolic process | |
| GO:0008967 | 1.9e-15 | phosphoglycolate phosphatase activity | ||
| GO:0003824 | 3.2e-12 | catalytic activity | ||
| KEGG pathway | ||||
| InterPro domain | [46-189] IPR023214 | 1.3e-31 | HAD-like domain | |
| [60-154] IPR006439 | 1.9e-15 | HAD-superfamily hydrolase, subfamily IA, variant 1 | ||
| [44-151] IPR005834 | 3.2e-12 | Haloacid dehalogenase-like hydrolase | ||
| Orthology group | MCL14657 | Multiple-copy universal gene | ||
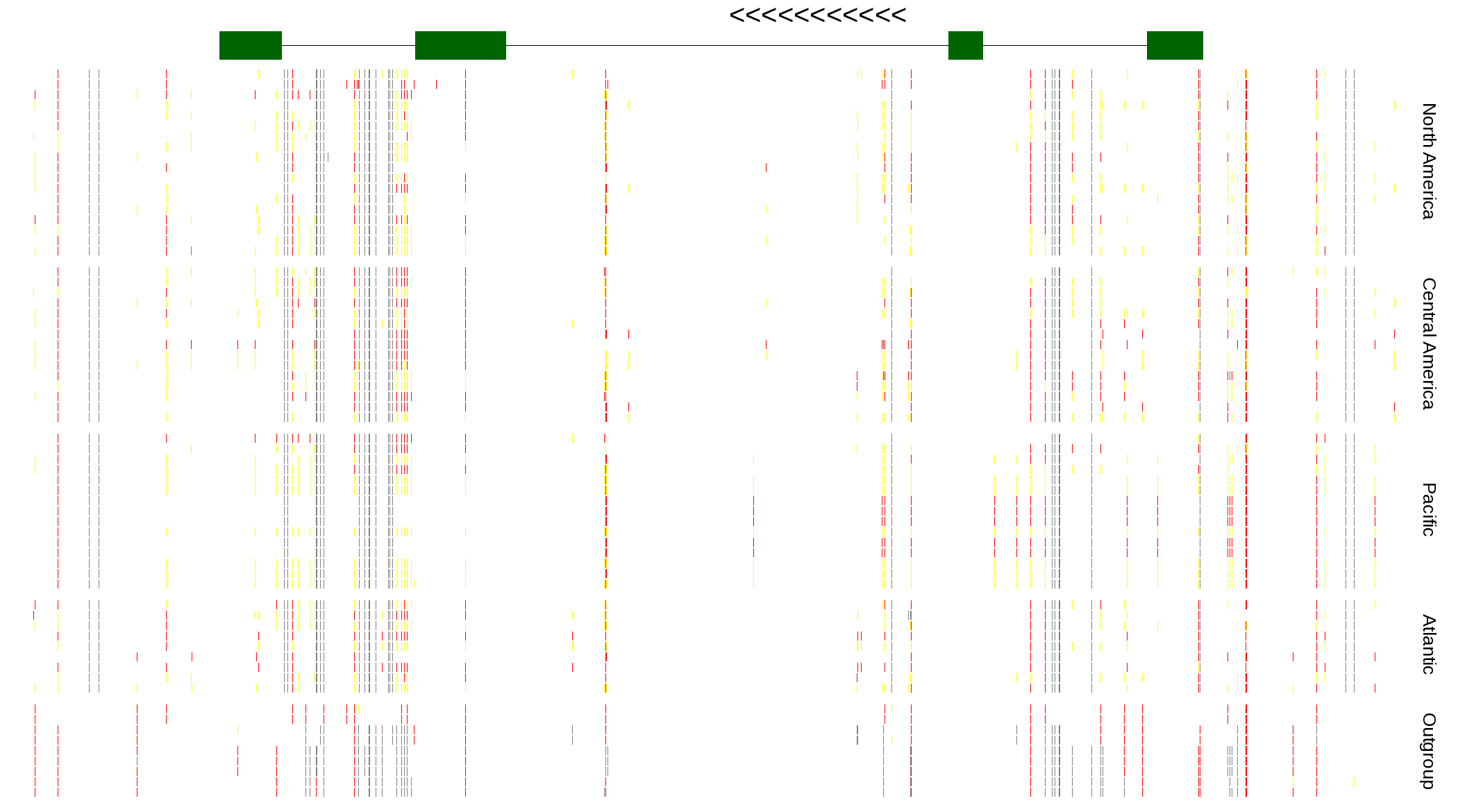
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204154-TA
ATGGATGCCGCTTCGGACAGTGCGTCTGCTTTCCTACAAGCATTCAGAGCAAAACCAGACGATGAATCCTACGCAATAGACGAATGGCCCACGCACTTATGGCGAAATTGTCTGCCTAAAAACTATAGGCATATTGCCAAAAATATATCAGCACAATGGTTAACATTAAGGTTTCAATATTTAGCACTGACGCCAGATGTGATACATCTTTTAGAATCTCTGAGGGAGGATTATCTTCTCGGTTTGATCACGAACGGGCCTTCGAGAGCACAATGGGAGAAGATCCAACGGCTGGATCTTGAGAAGTATTTCGACTGCGTGCTAGTCTCCGGGGATCATCCCTGGGAGAAACCAGATCAGAACATCTTCTTAGAAGCCTGCAAGTTACTCAATGTTGAAGCCAGAACTTGCATCATGGTTGGAGACAAACTCGAAACTGATATTAAGGGTGGTAAGGACGCTGAACTAGGAGGAACGGTGTGGATTCCTCTGCAGTATGATGAAATGGTATCAGATCTGCCAGATTTCAAAATTCAAAAGGTGACAGAGTTACCGGAAGTGCTGCCAAACTCACCAAAGCTAAAGAAGAGGGCTGCCTAA
>DPOGS204154-PA
MDAASDSASAFLQAFRAKPDDESYAIDEWPTHLWRNCLPKNYRHIAKNISAQWLTLRFQYLALTPDVIHLLESLREDYLLGLITNGPSRAQWEKIQRLDLEKYFDCVLVSGDHPWEKPDQNIFLEACKLLNVEARTCIMVGDKLETDIKGGKDAELGGTVWIPLQYDEMVSDLPDFKIQKVTELPEVLPNSPKLKKRAA-