| DPOGS204274 | ||
|---|---|---|
| Transcript | DPOGS204274-TA | 696 bp |
| Protein | DPOGS204274-PA | 231 aa |
| Genomic position | DPSCF300046 - 15617-17586 | |
| RNAseq coverage | 349x (Rank: top 33%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL003324 | 8e-121 | 87.07% | |
| Bombyx | BGIBMGA007544-TA | 8e-110 | 79.83% | |
| Drosophila | CG7785-PA | 3e-59 | 43.35% | |
| EBI UniRef50 | UniRef50_B0WY44 | 4e-72 | 54.98% | Putative uncharacterized protein n=2 Tax=Culicinae RepID=B0WY44_CULQU |
| NCBI RefSeq | XP_001862316.1 | 8e-73 | 54.98% | conserved hypothetical protein [Culex quinquefasciatus] |
| NCBI nr blastp | gi|170052644 | 1e-71 | 54.98% | conserved hypothetical protein [Culex quinquefasciatus] |
| NCBI nr blastx | gi|170052644 | 6e-71 | 54.98% | conserved hypothetical protein [Culex quinquefasciatus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 8.4e-14 | protein binding | |
| KEGG pathway | ||||
| InterPro domain | [4-229] IPR008985 | 7.3e-18 | Concanavalin A-like lectin/glucanase | |
| [62-207] IPR003877 | 8.4e-14 | SPla/RYanodine receptor SPRY | ||
| [61-214] IPR018355 | 3.7e-11 | SPla/RYanodine receptor subgroup | ||
| Orthology group | MCL13737 | Single-copy universal gene | ||
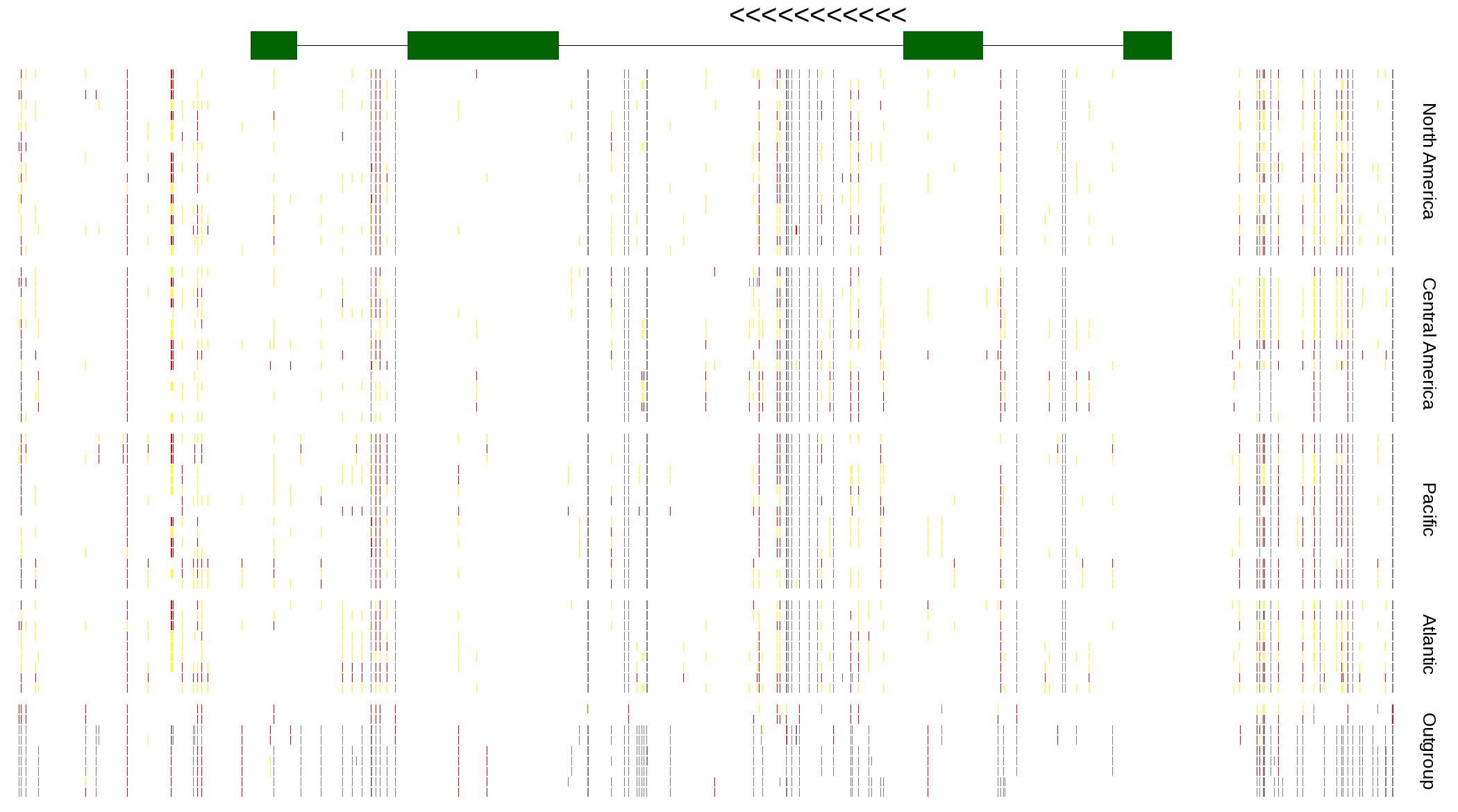
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204274-TA
ATGTTTTGCTGCTTAAAAAATTGCGTCAATGGCTTTTCTCTAACGCCAACTGTTCCTATACGTGTGAAGAGTAATCCAGTGCAGTTAGATTCTTTACATATGGGACATGAAATAGTCATAGTGAAAGGAGGCCAGCGTGTTTGTGGGTCCGGCTGTGCACTTGGCAATGCACCACTTGTTCAGAACAAAGCCTATTTTGAAGTAAAATTGCAGCAGGGTGGTGTGTGGGCAGTGGGACTTGCGACCAGGGAAGCAGACCTCAATAGAGTACATGGTGGTGTAGACAAAGAGTCTTGGTGTTTGAACAGTGACGGTACGGTTAGGCATGATAATGTTGAACTATACCATCTGAAACCAGCAAGTCAGGAGTCACCTACAGAGATCCTTGTCACAACTGGGTCGCCCAATCACACAGAGGATAAAGTTAACGACAAAATGGACAGTGAAAAAGCTAATGATAAGACAACAGCTATGCCCGGAGAGGGTGACGTAATTGGGGTAGCATATGATCATGTGGAGATGAACTTTTTCCTTAATGGAAAGAATATGGAAATTCCAGTCAGACAAATCAAGGGAACAGTGTACCCCGCTCTTTATGTTGATGACGGTGCAATTCTGGACATTATCCTTGACAACTTCAGATACCCACCACCGACGGGATATGAAAAGATCATGGTGGAGCAGTCACTCCTATAA
>DPOGS204274-PA
MFCCLKNCVNGFSLTPTVPIRVKSNPVQLDSLHMGHEIVIVKGGQRVCGSGCALGNAPLVQNKAYFEVKLQQGGVWAVGLATREADLNRVHGGVDKESWCLNSDGTVRHDNVELYHLKPASQESPTEILVTTGSPNHTEDKVNDKMDSEKANDKTTAMPGEGDVIGVAYDHVEMNFFLNGKNMEIPVRQIKGTVYPALYVDDGAILDIILDNFRYPPPTGYEKIMVEQSLL-