| DPOGS204507 | ||
|---|---|---|
| Transcript | DPOGS204507-TA | 432 bp |
| Protein | DPOGS204507-PA | 143 aa |
| Genomic position | DPSCF300002 + 1808306-1809033 | |
| RNAseq coverage | 144x (Rank: top 54%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006562 | 2e-40 | 65.55% | |
| Bombyx | BGIBMGA013511-TA | 3e-11 | 40.85% | |
| Drosophila | l(2)06225-PA | 2e-11 | 39.44% | |
| EBI UniRef50 | UniRef50_Q9VKM3 | 3e-09 | 39.44% | Lethal (2) 06225, isoform A n=32 Tax=Arthropoda RepID=Q9VKM3_DROME |
| NCBI RefSeq | NP_001165821.1 | 7e-11 | 37.50% | ATP synthase subunit g, mitochondrial [Nasonia vitripennis] |
| NCBI nr blastp | gi|340714767 | 4e-11 | 37.50% | PREDICTED: ATP synthase subunit g, mitochondrial-like [Bombus terrestris] |
| NCBI nr blastx | gi|340714767 | 8e-11 | 37.50% | PREDICTED: ATP synthase subunit g, mitochondrial-like [Bombus terrestris] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0015986 | 7.7e-16 | ATP synthesis coupled proton transport | |
| GO:0000276 | 7.7e-16 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o) | ||
| GO:0015078 | 7.7e-16 | hydrogen ion transmembrane transporter activity | ||
| KEGG pathway | nvi:100121208 | 2e-10 | ||
| K02140 (ATPeFG, ATP5L) | maps-> | Oxidative phosphorylation | ||
| InterPro domain | [63-143] IPR006808 | 7.7e-16 | ATPase, F0 complex, subunit G, mitochondrial | |
| Orthology group | MCL34551 | Lepidoptera specific | ||
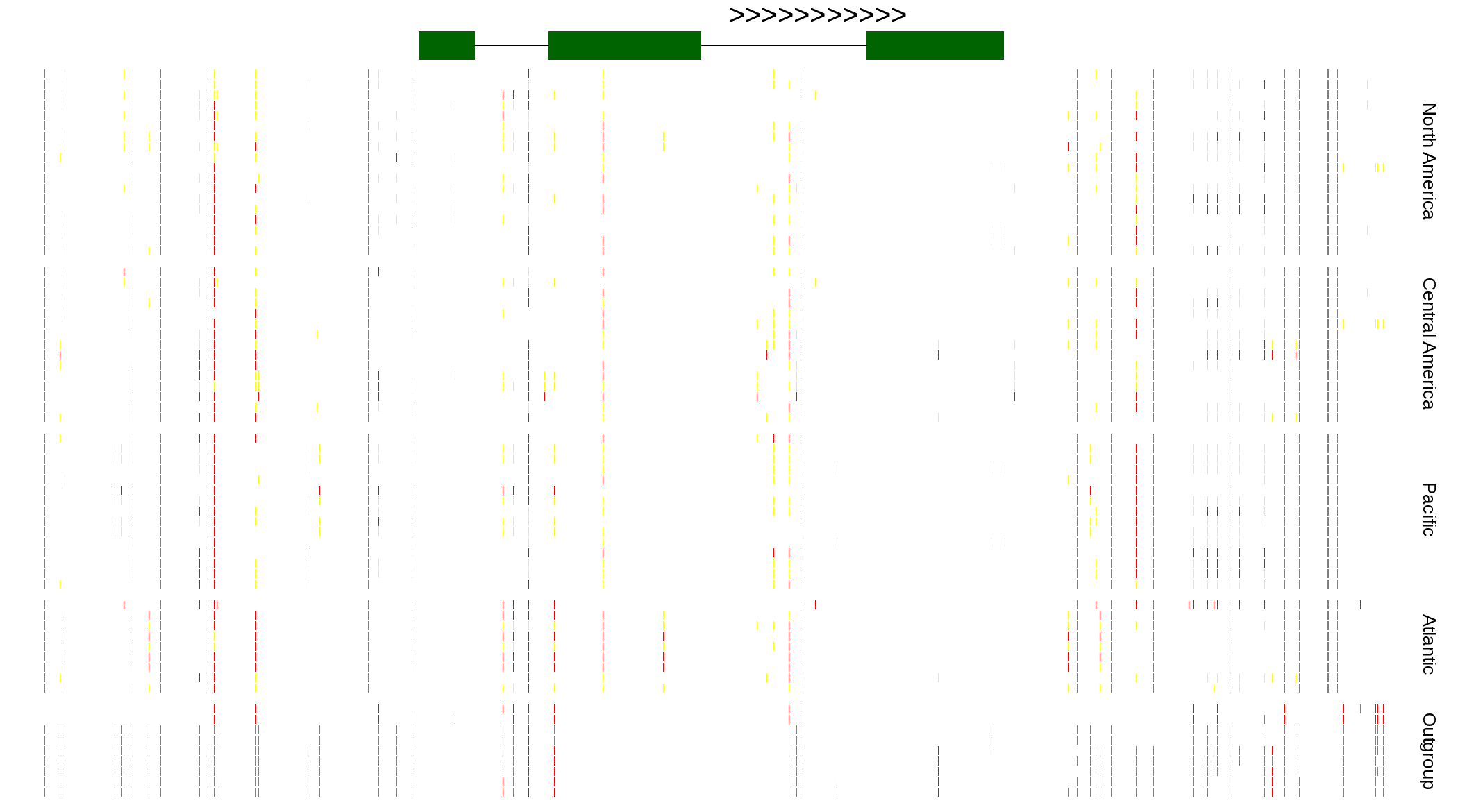
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204507-TA
ATGCAGATATTCCAATCAAGAAACATGACAACTAAAACGAAATCTGTTAAAGTGGGGATTCGATATTTACGTACAAGGTCGGCTTTGTTTTCGGAAATTGCCAATGTCTACAGACCAGCTTTGAAGCATCGTGAGGAAATATTGGAACTAATAAAAGAGAAATATTCAAAAGCATTGCAATCACGTTTTGCTGATAAAATGCGTACTGCTAAAGGGTTTTACAGATTAGAAATGTCTGTACCGAAAGTTGAAGAAATGAAGAGGATTCAAGAAGATTTGGCCTTAGTGAAGGATTTCATTAAAAATGAATGCTACAAGCAAATAACTGTAAAACAAGCTTGGTTACTCTTCCTCGTGGGATTGGAAATAGGGCTATGGTTCTTTCTAGGAGAAACCATTGGAAAATTTCATATTGTTGGATATAAAGTCTGA
>DPOGS204507-PA
MQIFQSRNMTTKTKSVKVGIRYLRTRSALFSEIANVYRPALKHREEILELIKEKYSKALQSRFADKMRTAKGFYRLEMSVPKVEEMKRIQEDLALVKDFIKNECYKQITVKQAWLLFLVGLEIGLWFFLGETIGKFHIVGYKV-