| DPOGS204665 | ||
|---|---|---|
| Transcript | DPOGS204665-TA | 486 bp |
| Protein | DPOGS204665-PA | 161 aa |
| Genomic position | DPSCF300170 - 383299-389471 | |
| RNAseq coverage | 218x (Rank: top 45%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL016249 | 4e-69 | 97.64% | |
| Bombyx | BGIBMGA007467-TA | 3e-16 | 100.00% | |
| Drosophila | Cdk5alpha-PA | 6e-67 | 82.61% | |
| EBI UniRef50 | UniRef50_Q17C39 | 2e-69 | 89.13% | Cyclin-dependent kinase 5 activator n=15 Tax=Bilateria RepID=Q17C39_AEDAE |
| NCBI RefSeq | XP_001649657.1 | 4e-70 | 89.13% | cyclin-dependent kinase 5 activator [Aedes aegypti] |
| NCBI nr blastp | gi|157107174 | 7e-69 | 89.13% | cyclin-dependent kinase 5 activator [Aedes aegypti] |
| NCBI nr blastx | gi|118788678 | 2e-65 | 89.13% | AGAP008533-PA [Anopheles gambiae str. PEST] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016533 | 1.5e-104 | cyclin-dependent protein kinase 5 holoenzyme complex | |
| GO:0016534 | 1.5e-104 | cyclin-dependent protein kinase 5 activator activity | ||
| KEGG pathway | aag:AaeL_AAEL004717 | 1e-69 | ||
| K11716 (CDK5R1) | maps-> | Alzheimer's disease | ||
| InterPro domain | [1-138] IPR004944 | 1.5e-104 | Cyclin-dependent kinase 5 activator | |
| [1-141] IPR013763 | 5.1e-76 | Cyclin-like | ||
| Orthology group | MCL11123 | Multiple-copy universal gene | ||
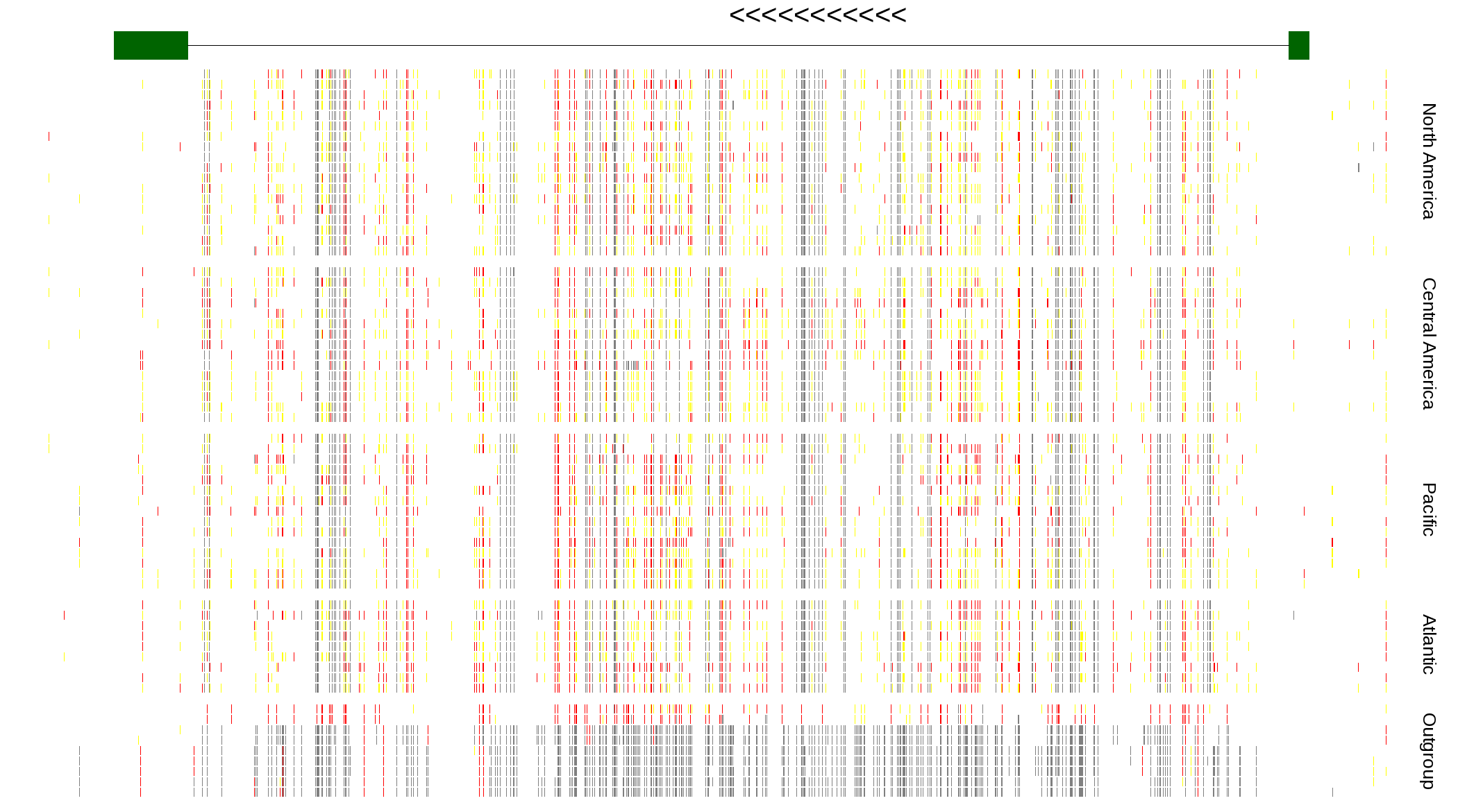
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204665-TA
ATGTTCCTCCACGCGCGCTGTCACCGGCTGCGTGACTTCCAGGCTGGAGACGCGGTCATGTGGCTGAGGACGGTAGACCGCTCGCTGCTCCTGCAAGGTTGGCAGGACGTGGCGTTCATCAATCCAGCTAACGTGGTCTTCGTGTATATGCTGGTTCGGGAGCTGGTGGACGGCGAGCGAATAGCTCGCCCCCAGGAACTACAAGCTGTAGTGCTCACCTGCCTCTACCTCTCTTACTCCTACATGGGCAACGAGATCTCCTATCCATTGAAACCCTTCCTCGTGGAGGATTCCAAGGATAAGTTTTGGGATCGCTGTCTGCTCATCGTTGACAGGCTATCCTTCAATATGTTGAGAATCAACTCGGAGCCTGGGTTCTTCACTGAAGTGTTCACGGAGTTGAAAGCGTGTGGAACTGTGGATAGCCCCCCTCCCGCACCCCTCCATCCAGCGCCTGCGCCCCCTCACGCCTTGACGGCGGCCTGA
>DPOGS204665-PA
MFLHARCHRLRDFQAGDAVMWLRTVDRSLLLQGWQDVAFINPANVVFVYMLVRELVDGERIARPQELQAVVLTCLYLSYSYMGNEISYPLKPFLVEDSKDKFWDRCLLIVDRLSFNMLRINSEPGFFTEVFTELKACGTVDSPPPAPLHPAPAPPHALTAA-