| DPOGS205993 | ||
|---|---|---|
| Transcript | DPOGS205993-TA | 570 bp |
| Protein | DPOGS205993-PA | 189 aa |
| Genomic position | DPSCF300164 + 322743-326743 | |
| RNAseq coverage | 942x (Rank: top 14%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL005154 | 6e-71 | 96.06% | |
| Bombyx | BGIBMGA009419-TA | 4e-95 | 84.04% | |
| Drosophila | CG5941-PA | 5e-78 | 71.91% | |
| EBI UniRef50 | UniRef50_G6DBG4 | 1e-101 | 100.00% | Mct-1 protein n=32 Tax=Eukaryota RepID=G6DBG4_DANPL |
| NCBI RefSeq | XP_001658619.1 | 8e-90 | 85.96% | mct-1 protein [Aedes aegypti] |
| NCBI nr blastp | gi|307187668 | 9e-91 | 87.08% | Malignant T cell amplified sequence 1 [Camponotus floridanus] |
| NCBI nr blastx | gi|307187668 | 4e-88 | 87.08% | Malignant T cell amplified sequence 1 [Camponotus floridanus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003723 | 4.7e-23 | RNA binding | |
| KEGG pathway | ||||
| InterPro domain | [1-189] IPR016437 | 1.7e-124 | Translation-associated RNA-binding, predicted | |
| [74-180] IPR004521 | 4.7e-23 | Uncharacterised domain CHP00451 | ||
| [99-187] IPR015947 | 5.1e-21 | PUA-like domain | ||
| [103-178] IPR002478 | 1.7e-17 | Pseudouridine synthase/archaeosine transglycosylase | ||
| Orthology group | MCL12521 | Single-copy universal gene | ||
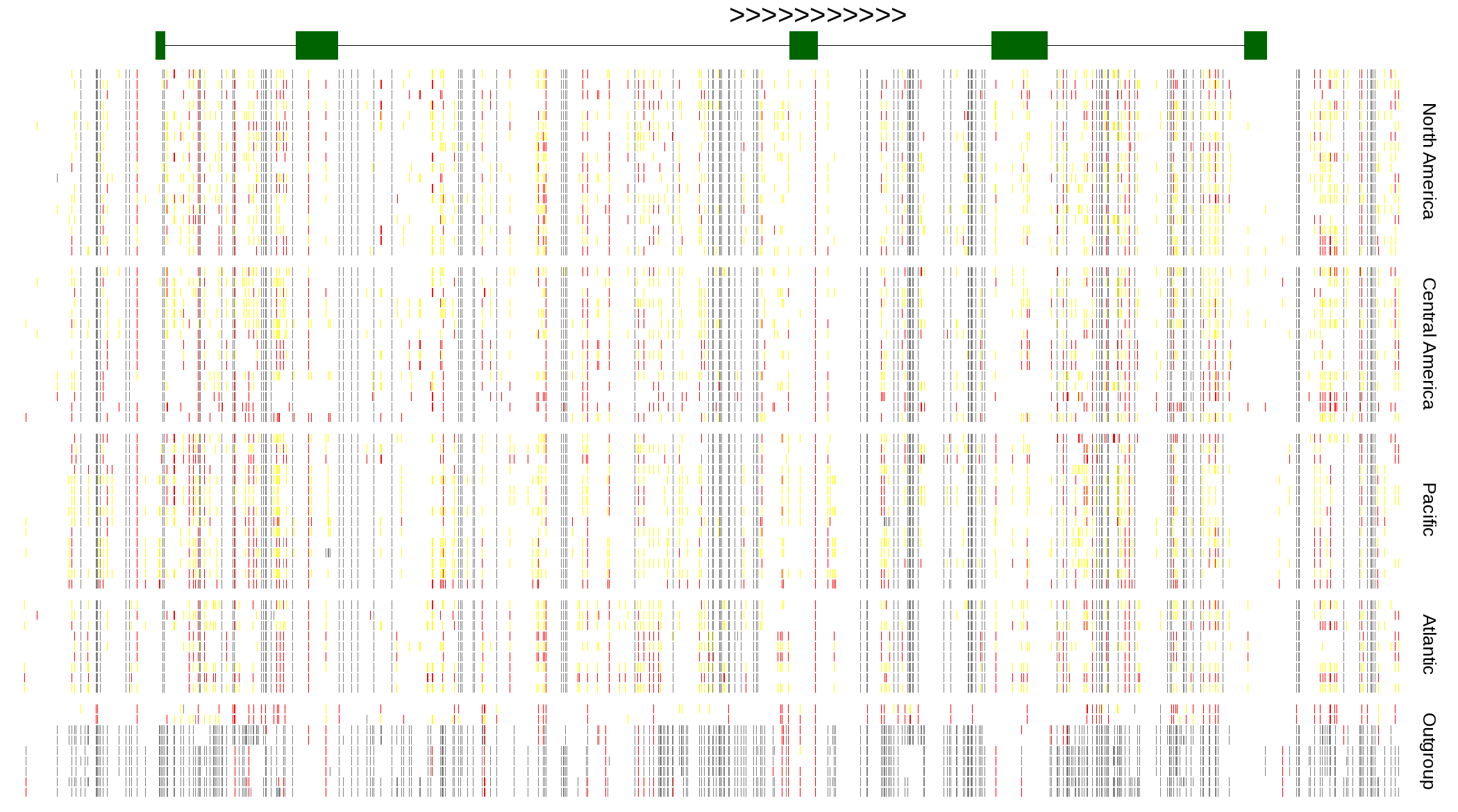
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS205993-TA
ATGTACAAAAAAGTAATGAAGTTTGGGATAATATTTGATGAAAAGGAAAGCATTTCAGGCGTTCAGCAGCTGAAGTCCTCGGTACAGAAGGGCATCAGATCCCGTCTGTTGGAACTGTACCCACATCTGGAGAACCACATTGACCAAGTACTACCCAAGAAGGATACATTTAGGATTGTCAAATGTCATGACCACATCGAGATCATGGTGAACAGTGCCGGGGAATTGCTGTTCTTCAGACACAGAGAAGGTCCTTGGATGCCAACCTTGAGGCTGCTACATAAATATCCATTCTTCTTACCAATGCAGCAAGTCGACAAAGGTGCTATAAGATTCGTTCTTAGCGGTGCTAATATAATGTGCCCCGGACTGACTTCACCTGGAGCACGTATGTCGAGTGTAGTGAAAGGCAGTGTTGTAGCTGTTATGGCCGAGGGCAAACAGCATGCGTTGGCTGTTGGCATAACTTCACTATCAACTGATGATATAGCGAAAGTCAACAAGGGCGTCGGCATAGAGAACTGTCACTATTTGAATGATGGCTTATGGCAGATGAAACCAGTGAAGTGA
>DPOGS205993-PA
MYKKVMKFGIIFDEKESISGVQQLKSSVQKGIRSRLLELYPHLENHIDQVLPKKDTFRIVKCHDHIEIMVNSAGELLFFRHREGPWMPTLRLLHKYPFFLPMQQVDKGAIRFVLSGANIMCPGLTSPGARMSSVVKGSVVAVMAEGKQHALAVGITSLSTDDIAKVNKGVGIENCHYLNDGLWQMKPVK-