| DPOGS206167 | ||
|---|---|---|
| Transcript | DPOGS206167-TA | 333 bp |
| Protein | DPOGS206167-PA | 110 aa |
| Genomic position | DPSCF300434 - 4395-4727 | |
| RNAseq coverage | 0x (Rank: top 97%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | % | |||
| Bombyx | % | |||
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_Q53H47 | 1e-21 | 47.71% | Histone-lysine N-methyltransferase SETMAR n=98 Tax=Bilateria RepID=SETMR_HUMAN |
| NCBI RefSeq | XP_001119968.1 | 5e-19 | 39.81% | PREDICTED: similar to SET domain and mariner transposase fusion [Apis mellifera] |
| NCBI nr blastp | gi|85679599 | 1e-21 | 47.71% | SETMAR [Homo sapiens] |
| NCBI nr blastx | gi|384943140 | 2e-20 | 47.71% | histone-lysine N-methyltransferase SETMAR [Macaca mulatta] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | mcc:703880 | 5e-22 | ||
| K11433 (SETMAR) | maps-> | Lysine degradation | ||
| Orthology group | MCL22623 | Insect specific | ||
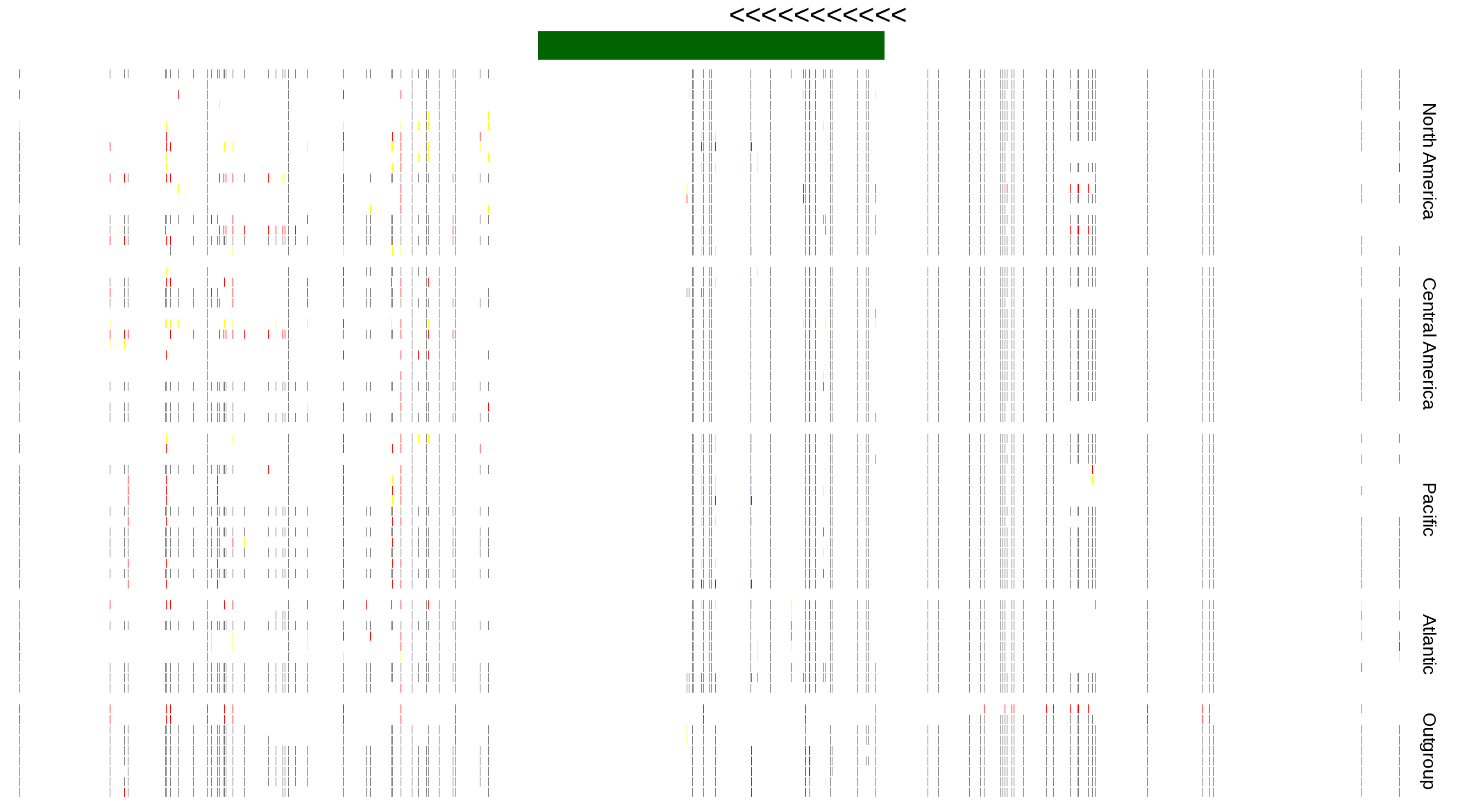
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206167-TA
ATGGCCGTATCAAATGCAATAATTCGTGCTTGTTTACTTTACGAATTCAAACTTGGCAGTAAAGCCGCAGAGGCTGGTCGTAAGATATGCACTGCTTTAAGGAAGGGTGCTGTAGCTGAACGAACGAGCCAGAAGTGGTTTAAAAAATTTGTATCTGGAAATGAAAGCCTGGATGATGAACCTCGCTCCGGAAGACCATCGACGATTAACAACGAAGATATCAAGTTGGCAATCGAGCAGGATTCAAGTCAGACATGCCAAGCTCTTGCCTTGCAATTCAATGTCAGCGACGAAACTATTCGCATGCATTTGCACCAGCTTGGGAAACGATAG
>DPOGS206167-PA
MAVSNAIIRACLLYEFKLGSKAAEAGRKICTALRKGAVAERTSQKWFKKFVSGNESLDDEPRSGRPSTINNEDIKLAIEQDSSQTCQALALQFNVSDETIRMHLHQLGKR-