| DPOGS206479 | ||
|---|---|---|
| Transcript | DPOGS206479-TA | 660 bp |
| Protein | DPOGS206479-PA | 219 aa |
| Genomic position | DPSCF300070 + 779622-780281 | |
| RNAseq coverage | 5x (Rank: top 88%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012710 | 2e-95 | 73.73% | |
| Bombyx | BGIBMGA005927-TA | 2e-53 | 60.13% | |
| Drosophila | CG1675-PA | 4e-54 | 45.41% | |
| EBI UniRef50 | UniRef50_E2AC78 | 2e-61 | 53.05% | Methyltransferase-like protein 11A n=10 Tax=Pancrustacea RepID=E2AC78_CAMFO |
| NCBI RefSeq | XP_623109.1 | 1e-64 | 52.75% | PREDICTED: similar to CG1675-PA isoform 1 [Apis mellifera] |
| NCBI nr blastp | gi|66562955 | 2e-63 | 52.75% | PREDICTED: alpha N-terminal protein methyltransferase 1A-like isoform 1 [Apis mellifera] |
| NCBI nr blastx | gi|66562955 | 4e-61 | 52.75% | PREDICTED: alpha N-terminal protein methyltransferase 1A-like isoform 1 [Apis mellifera] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0008168 | 5.4e-103 | methyltransferase activity | |
| KEGG pathway | ||||
| InterPro domain | [1-219] IPR008576 | 5.4e-103 | Protein of unknown function DUF858, methyltransferase-like | |
| Orthology group | MCL13308 | Single-copy universal gene | ||
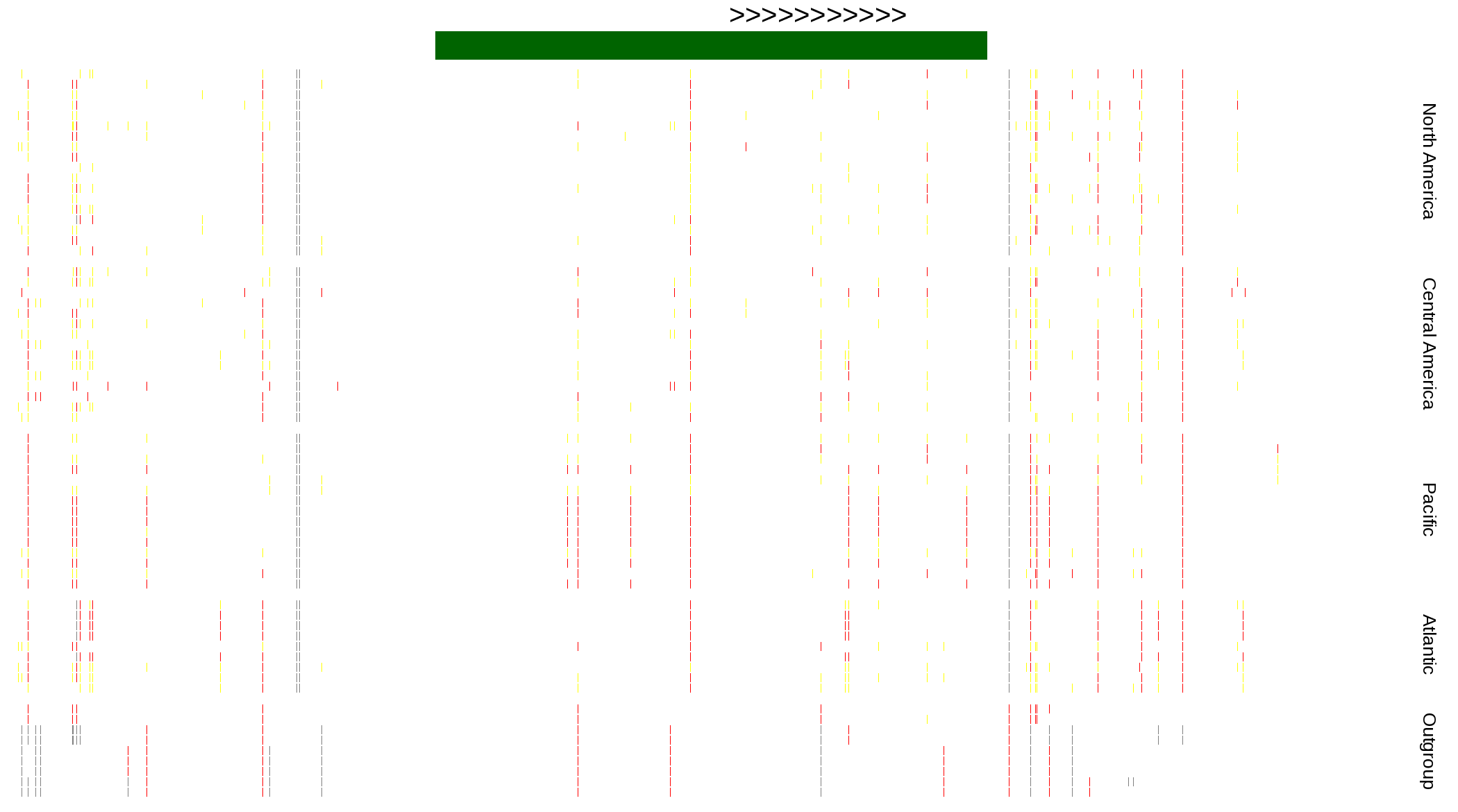
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206479-TA
ATGACGGAGACAACATTTTATAAGAAAGCCGCAAAATACTGGGCTAATGTACCTGCAACAATAGACGGAGTGCTCGGCGGCTTCGGCCACATTACGGACATTGACATCGAGGGGTCCAAGAAATTTTTAAATTACATATTGTCACTAGAGCAACCACCGAAAACCAATTTGGCCTTGGACTGCGGAGCCGGTATTGGAAGAGTCAGCAAGAATCTTTTGATGCATTACTTCGTGAAGGTCGATTTAGTGGAACAAGACGAAAAGTTCATTACCACGGCAAAGCAATTGCTTGGCGAAAACAATGCTAAATTGGGAACGTTGTACCAAATCGGGCTTCAACATCTTAAGCTACAAAAAAAATACGACATGATTTGGTGCCAGTGGGTGCTTGGGCATTTAAATGACTACGATTTAATTACATTCTTGGAAAGATGTAATCAAGCGTTGGCTGAAAATGGCGTGATTATTGTAAAAGAAAACATAGCACCATCAGACGAAATCGAATACGATGATGAAGATTCCTCTGTAACTCGTCCATATAAATTAATGGAGAAAATATTTGAGGATGCAAATCTTCGTGTCATCGCCACTGACATACAAAGTGGTTTCCCTGATGATATTTATCCCGTTCATATGTTTGCATTAGCGCCGTTTCAATAA
>DPOGS206479-PA
MTETTFYKKAAKYWANVPATIDGVLGGFGHITDIDIEGSKKFLNYILSLEQPPKTNLALDCGAGIGRVSKNLLMHYFVKVDLVEQDEKFITTAKQLLGENNAKLGTLYQIGLQHLKLQKKYDMIWCQWVLGHLNDYDLITFLERCNQALAENGVIIVKENIAPSDEIEYDDEDSSVTRPYKLMEKIFEDANLRVIATDIQSGFPDDIYPVHMFALAPFQ-