| DPOGS206800 | ||
|---|---|---|
| Transcript | DPOGS206800-TA | 582 bp |
| Protein | DPOGS206800-PA | 193 aa |
| Genomic position | DPSCF300001 - 4394391-4417143 | |
| RNAseq coverage | 3200x (Rank: top 4%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL009249 | 4e-87 | 81.38% | |
| Bombyx | BGIBMGA000619-TA | 9e-73 | 87.84% | |
| Drosophila | Fkbp13-PD | 1e-72 | 70.98% | |
| EBI UniRef50 | UniRef50_Q6NL72 | 8e-70 | 69.43% | RE40519p n=20 Tax=Opisthokonta RepID=Q6NL72_DROME |
| NCBI RefSeq | NP_001108408.1 | 5e-85 | 79.79% | FK506-binding protein [Bombyx mori] |
| NCBI nr blastp | gi|169234934 | 8e-84 | 79.79% | FK506-binding protein precursor [Bombyx mori] |
| NCBI nr blastx | gi|169234934 | 2e-83 | 79.79% | FK506-binding protein precursor [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006457 | 1.3e-31 | protein folding | |
| GO:0005509 | 3.3e-11 | calcium ion binding | ||
| KEGG pathway | ||||
| InterPro domain | [1-168] IPR023566 | 7.4e-72 | Peptidyl-prolyl cis-trans isomerase, FKBP-type | |
| [2-84] IPR001179 | 1.3e-31 | Peptidyl-prolyl cis-trans isomerase, FKBP-type, domain | ||
| [108-185] IPR011992 | 3.3e-11 | EF-hand-like domain | ||
| Orthology group | MCL14403 | Single-copy universal gene | ||
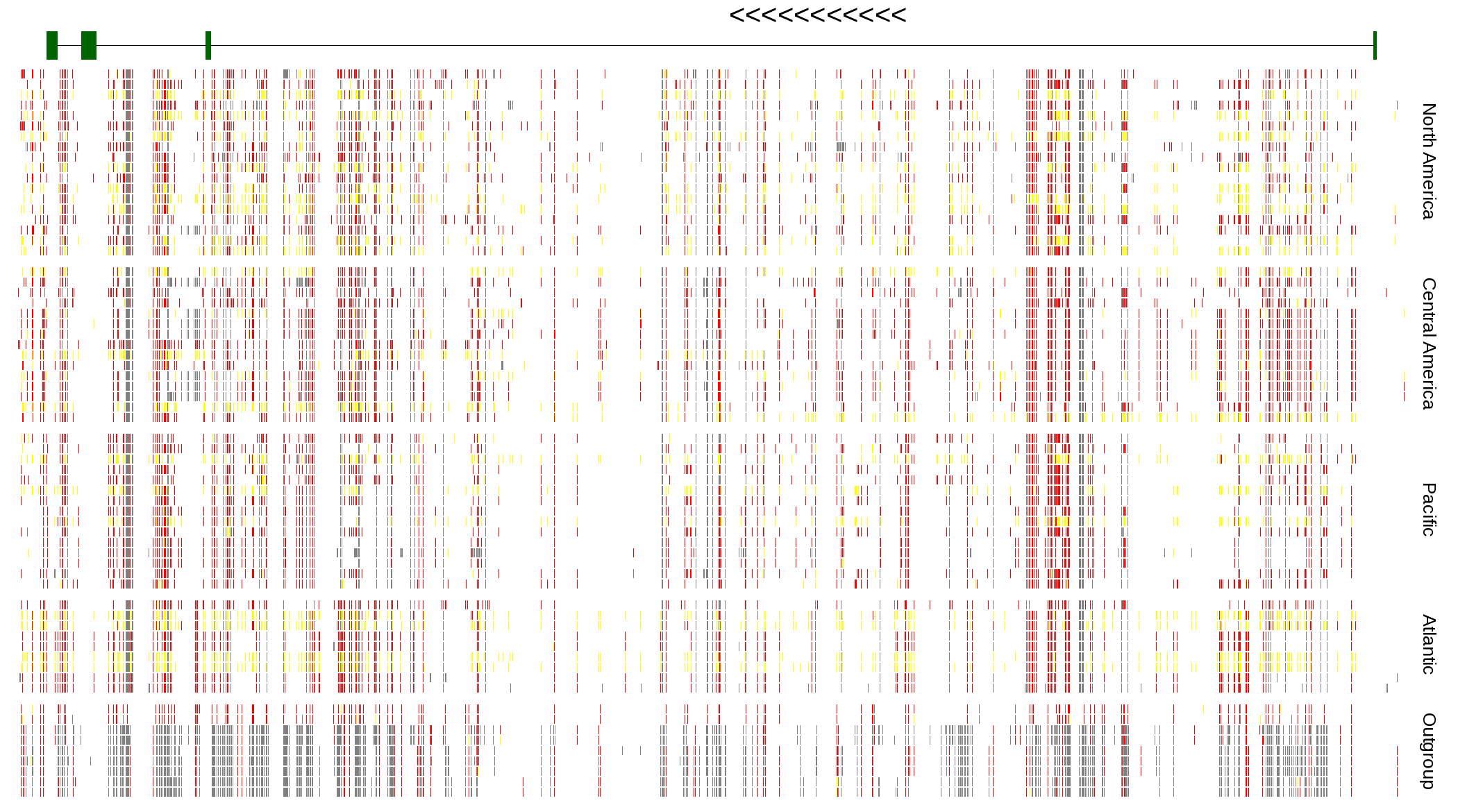
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206800-TA
ATGCTCACTATGCACTACACCGGCACGCTCGAGAACGGTCATAAGTTCGACTCCAGCTACGACCGTGATCAACCGTTTACCTTCCAGCTGGGAGTGGGTCAGGTCATCAAGGGATGGGACCAAGGATTAGTCGACATGTGTGTTGGTGAAAAACGTAAACTCGTTATTCCGTCATCCCTGGGCTACGGTGATCGCGGAGCTGGTAATGTCATCCCTGGAGGCGCAACACTCTTCTTTGATGTAGAACTGATCAACATCGGCGACACACCGCCCACAACCAACGTGTTCAAAGAAATCGACGCCGACAAAGACAACATGCTATCACGAGAGGAAGTGAGTATAGAAATAGTGTTCCGGGCCATGGACACGGACGGAGATAGCGAACTGTCTAGGGAAGAGGTGAGTGACTACCTGAAGAAGCAGATGGTGCCACAGGATGGTAGTGAGATGAGTGAAGATGTCAAGCAGATGTTGGAAAGTCACGACAAACTGGTAGAGGAGATCTTCCAACACGAAGACAAGGACAAGAACGGATTCATCAGCCATGAGGAGTTCTCTGGTCCTAAGCACGACGAACTCTAA
>DPOGS206800-PA
MLTMHYTGTLENGHKFDSSYDRDQPFTFQLGVGQVIKGWDQGLVDMCVGEKRKLVIPSSLGYGDRGAGNVIPGGATLFFDVELINIGDTPPTTNVFKEIDADKDNMLSREEVSIEIVFRAMDTDGDSELSREEVSDYLKKQMVPQDGSEMSEDVKQMLESHDKLVEEIFQHEDKDKNGFISHEEFSGPKHDEL-