| DPOGS206884 | ||
|---|---|---|
| Transcript | DPOGS206884-TA | 591 bp |
| Protein | DPOGS206884-PA | 196 aa |
| Genomic position | DPSCF300001 - 2041133-2041723 | |
| RNAseq coverage | 39x (Rank: top 73%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006878 | 5e-80 | 77.27% | |
| Bombyx | BGIBMGA012841-TA | 2e-81 | 72.34% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_UPI000203A21D | 1e-22 | 38.89% | UPI000203A21D related cluster n=1 Tax=unknown RepID=UPI000203A21D |
| NCBI RefSeq | XP_002738372.1 | 2e-22 | 37.32% | PREDICTED: methyl-CpG binding domain protein 4-like [Saccoglossus kowalevskii] |
| NCBI nr blastp | gi|327280386 | 5e-22 | 38.89% | PREDICTED: methyl-CpG-binding domain protein 4-like [Anolis carolinensis] |
| NCBI nr blastx | gi|328712829 | 4e-22 | 43.85% | PREDICTED: methyl-CpG-binding domain protein 4-like [Acyrthosiphon pisum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006281 | 1.2e-19 | DNA repair | |
| GO:0003824 | 1.2e-19 | catalytic activity | ||
| KEGG pathway | xtr:733528 | 3e-20 | ||
| K10801 (MBD4) | maps-> | Base excision repair | ||
| InterPro domain | [22-171] IPR011257 | 1.2e-19 | DNA glycosylase | |
| Orthology group | MCL19609 | Patchy | ||
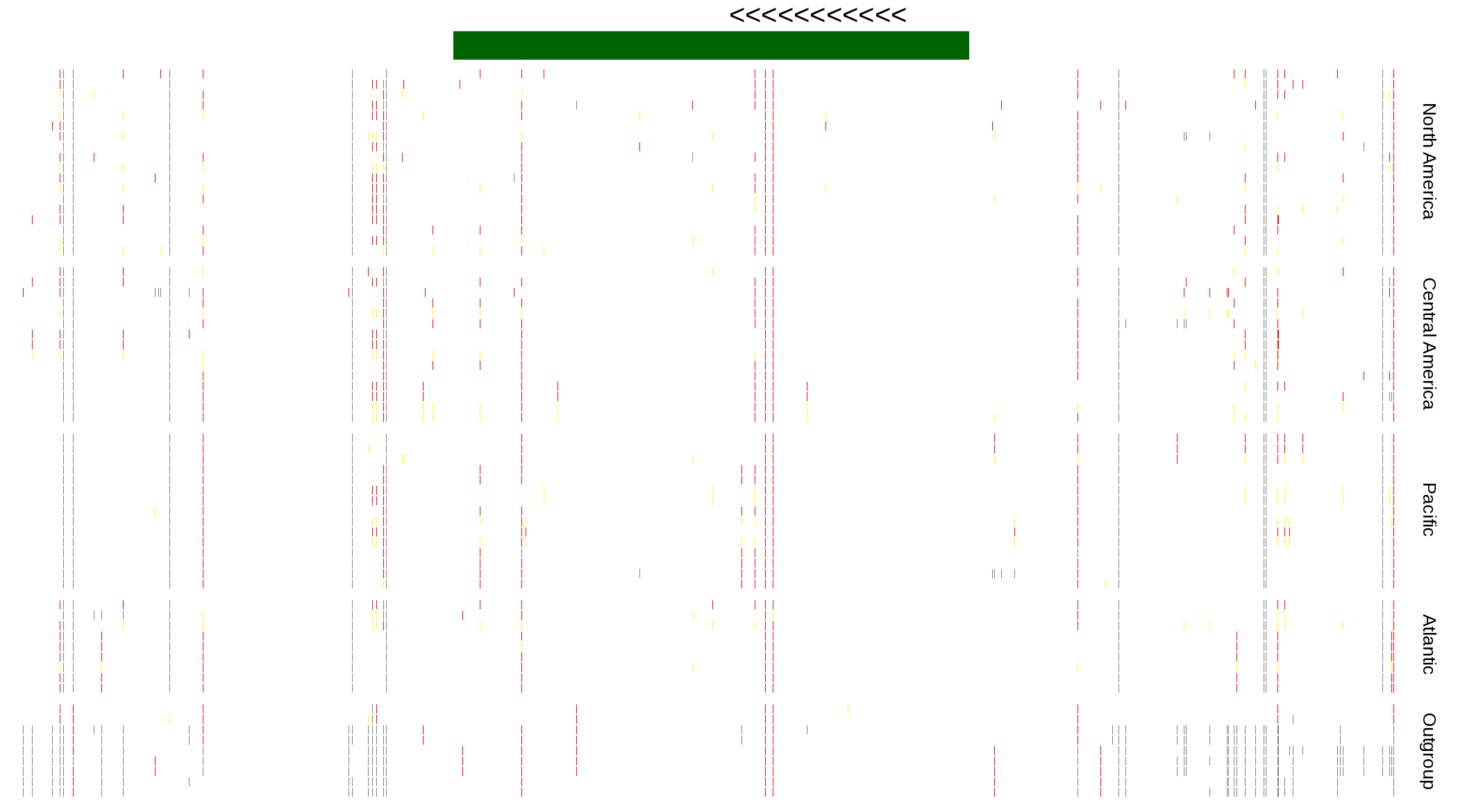
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206884-TA
ATGAATTTGGAGAACGACTTGAGTTTATCACAATTAACAATAGAAGAGCCGGACCCATTAAACATACCGCCTTTCTTCAATCTCACACCTCGCTTAATGCCCGAATCACCTCACTATATAATAGAGGAGGAATTCTCTCTCAACCCCTGGGCAATGTTAGTAGCAACTATATTCCTAACAAAGACGTCCGGCAAAACAGCTAGGCCGTATATAAAGAGTTTTTTTACGGACTACCCGACACCTTATCAAGTTTTGGATGAAACTCCATCGTCATTAGAGAGGTTCTTCGAGAACTTAGGATTAAAAAAACGTGGTAATATGATTTGGAAGCTGAGTTACCAGTTCGTGTCTGGTAAATGGCGGCGAGCTAGCGATCTCTGTGGAATCGGGAAGTATGGGGAGGACGCTTATAGGATCTTTTGCTTAGGTCACACGGATGTAAACCCTGACGATAGATATTTAAAGCTCTATTTAGATTGGCTGCAGTGTCACACTGAGTTCATAAAAGACAGGAGCGTAACTGACAGCGAAAACCTATTACAAGATCCGGTTCTGAAATATTATAGAATTACTTTGAAAAGTAATGTATAA
>DPOGS206884-PA
MNLENDLSLSQLTIEEPDPLNIPPFFNLTPRLMPESPHYIIEEEFSLNPWAMLVATIFLTKTSGKTARPYIKSFFTDYPTPYQVLDETPSSLERFFENLGLKKRGNMIWKLSYQFVSGKWRRASDLCGIGKYGEDAYRIFCLGHTDVNPDDRYLKLYLDWLQCHTEFIKDRSVTDSENLLQDPVLKYYRITLKSNV-