| DPOGS206970 | ||
|---|---|---|
| Transcript | DPOGS206970-TA | 633 bp |
| Protein | DPOGS206970-PA | 210 aa |
| Genomic position | DPSCF300001 + 189623-191440 | |
| RNAseq coverage | 91x (Rank: top 63%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL014354 | 4e-113 | 90.48% | |
| Bombyx | BGIBMGA012874-TA | 4e-56 | 77.87% | |
| Drosophila | CG4433-PA | 1e-61 | 48.71% | |
| EBI UniRef50 | UniRef50_B4PP52 | 2e-63 | 54.03% | GE25096 n=8 Tax=Endopterygota RepID=B4PP52_DROYA |
| NCBI RefSeq | XP_001979470.1 | 1e-64 | 54.03% | GG15675 [Drosophila erecta] |
| NCBI nr blastp | gi|386766122 | 1e-63 | 54.03% | CG4433, isoform C [Drosophila melanogaster] |
| NCBI nr blastx | gi|386766122 | 4e-62 | 54.03% | CG4433, isoform C [Drosophila melanogaster] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | der:Dere_GG15675 | 4e-64 | ||
| K03434 (E3.5.1.89) | maps-> | Glycosylphosphatidylinositol(GPI)-anchor biosynthesis | ||
| InterPro domain | [1-210] IPR003737 | 7.2e-82 | N-acetylglucosaminyl phosphatidylinositol deacetylase | |
| [1-195] IPR024078 | 5.3e-48 | Putative deacetylase LmbE-like domain | ||
| Orthology group | MCL12920 | Single-copy universal gene | ||
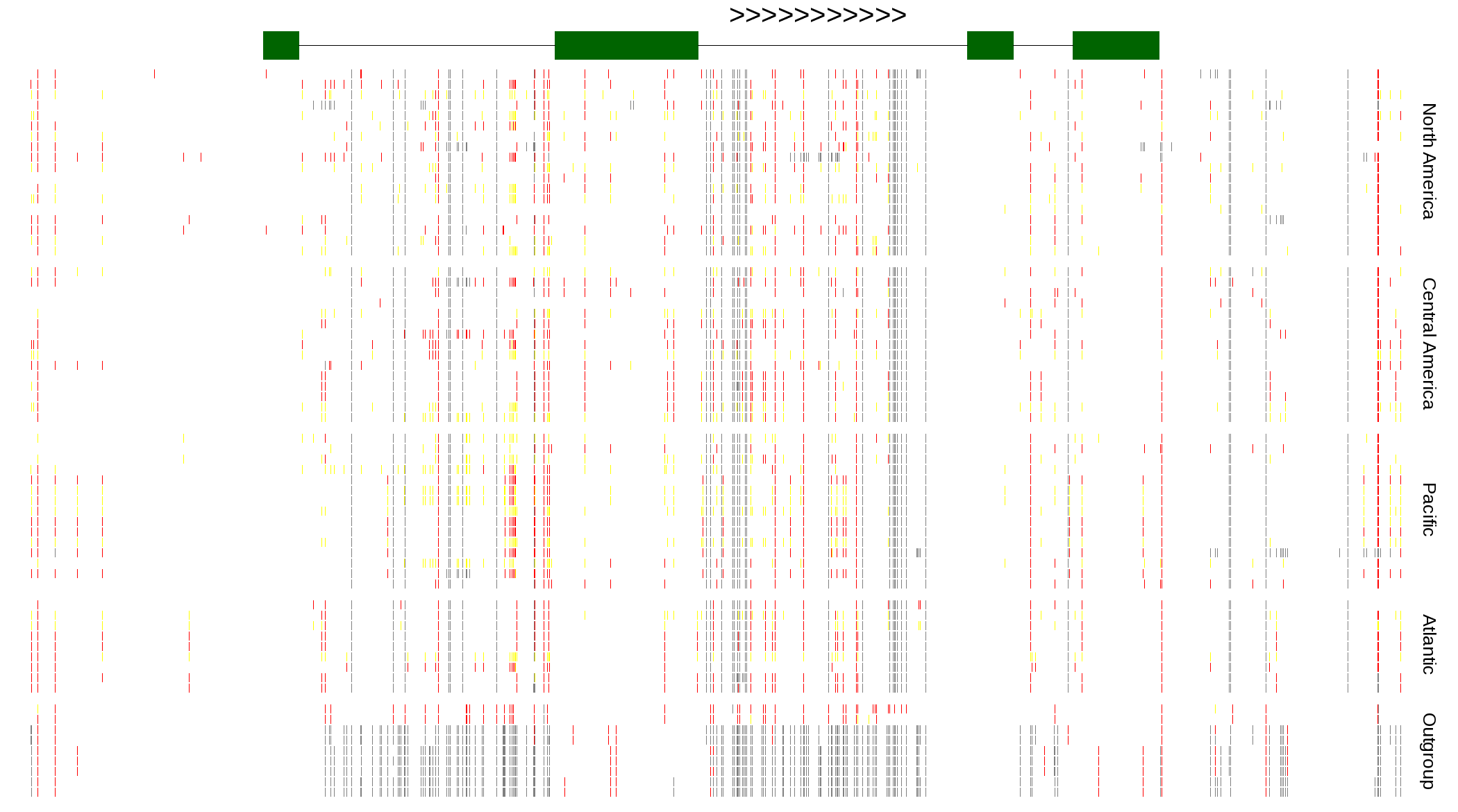
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206970-TA
ATGTTCTTTGGTCCAACAATATTTAGATTGTGCGAGCAAGGTGCTGAAGTCTATCTGCTGTGTCTATCCAATGGTAATTATGAGGGCAAAGGCGGCGAAAGACGCAAGGAGCTGTGGAATGCCTGCCGTGAGCTTGGTGTACCTGATAGCAATATATGTCTCATTATGGACACAAGATTGCCGGACGACCCTAAAGCGCAATGGCCAGTTCCTGTTGTAGCTAGACTCATACAACACAAGTTAGAGGCTTTGGATATAGATACATTAGTGACATTTGATCGAGGAGGTGTTTCATCACACCCAAACCACTCAGCTGCCTTTTATGCTGTAGCTTATATGTTTGTTGAAAAGAATATGCCTTCAAAATGTACTTTTTATACACTGGATTCTGTAAATATACTGAGAAAGTACTTGGGGTTCCTAGATCTTCCACTGAGCTTTGTTCTATCTTCCAAAAGGTACTTCCTTCGCTGGACTGAGAGTCGTCGCGTGACACGAGCGATGAAGCTTCACAAGAGCCAGATGGTGTGGTTCAGATATCTGTATGTGATGTTTTCTAGGTACATGGTCATCAATACACTTAGGAAGATTAATCTCGCCGATATAGAGCTTGAACTAGAAGTAGATGACTAG
>DPOGS206970-PA
MFFGPTIFRLCEQGAEVYLLCLSNGNYEGKGGERRKELWNACRELGVPDSNICLIMDTRLPDDPKAQWPVPVVARLIQHKLEALDIDTLVTFDRGGVSSHPNHSAAFYAVAYMFVEKNMPSKCTFYTLDSVNILRKYLGFLDLPLSFVLSSKRYFLRWTESRRVTRAMKLHKSQMVWFRYLYVMFSRYMVINTLRKINLADIELELEVDD-