| DPOGS206980 | ||
|---|---|---|
| Transcript | DPOGS206980-TA | 414 bp |
| Protein | DPOGS206980-PA | 137 aa |
| Genomic position | DPSCF300001 + 420725-426677 | |
| RNAseq coverage | 2324x (Rank: top 5%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL002124 | 2e-54 | 67.88% | |
| Bombyx | BGIBMGA012942-TA | 7e-47 | 66.18% | |
| Drosophila | fabp-PC | 9e-18 | 47.79% | |
| EBI UniRef50 | UniRef50_E2BJH6 | 9e-20 | 43.28% | Myelin P2 protein n=18 Tax=Coelomata RepID=E2BJH6_HARSA |
| NCBI RefSeq | NP_001155787.1 | 2e-23 | 43.80% | hypothetical protein LOC100168295 [Acyrthosiphon pisum] |
| NCBI nr blastp | gi|383853580 | 4e-23 | 45.59% | PREDICTED: myelin P2 protein-like [Megachile rotundata] |
| NCBI nr blastx | gi|383853580 | 9e-23 | 45.59% | PREDICTED: myelin P2 protein-like [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005488 | 2.6e-33 | binding | |
| GO:0008289 | 1e-06 | lipid binding | ||
| GO:0006810 | 1e-06 | transport | ||
| GO:0005215 | 1e-06 | transporter activity | ||
| KEGG pathway | cfa:478156 | 1e-14 | ||
| K08752 (FABP3) | maps-> | PPAR signaling pathway | ||
| InterPro domain | [5-134] IPR012674 | 2.6e-33 | Calycin | |
| [2-135] IPR011038 | 2.6e-33 | Calycin-like | ||
| [8-117] IPR000566 | 1.6e-08 | Lipocalin/cytosolic fatty-acid binding protein domain | ||
| [5-27] IPR000463 | 1e-06 | Cytosolic fatty-acid binding | ||
| Orthology group | MCL26023 | Lepidoptera specific | ||
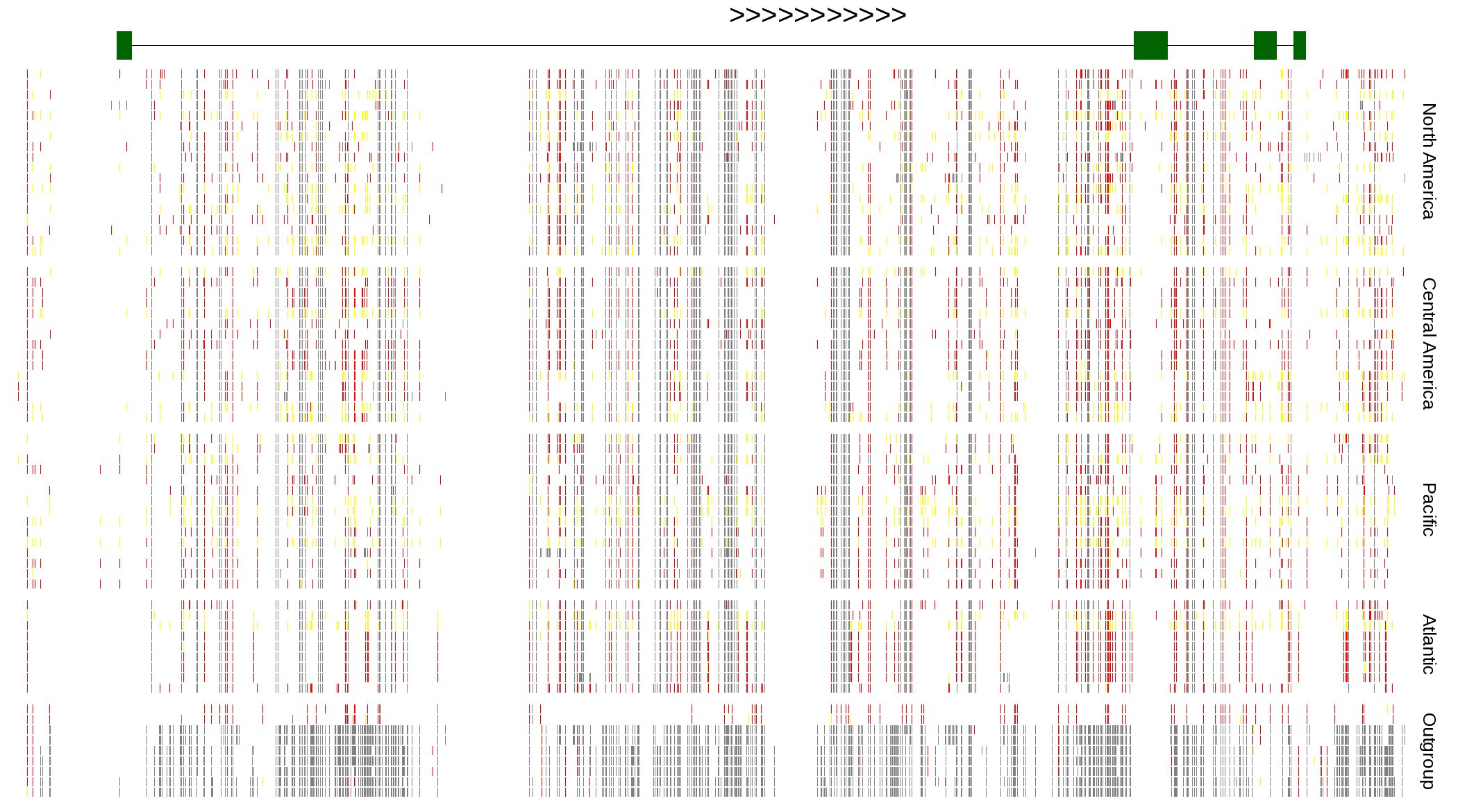
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206980-TA
ATGGAAAAATATCTTGGAAAAAAATATAAATTGAGATCATCGCATAACTTTGATGAATATCTCAAATTCATTGAAGTCGGCCTGTTATCTCGGAAGCTGGTGACTTCACTGTCCCCAATCAGTGTGCTGACCAAAAACGAAGATGGCTCATACTCTCTTACAATGATAACTCCGATACGGAAGGTTGTCATTACATTTCAACTCGGAGTGGAATTCTCAGAGGATAGACCAGATGGTATCAAAGTCAAGTCAACTATGCATCTTGATGGAGATAAGTTAATACAGACACAAATTGAAAGTAATGGTAGGAAGTCAACGCACGTGAGAGAGTTTACAGACAAGCTTTTAACAGTGACAACAACTGCTGAAGGTTGGGACGGACAATGCATTAGAATATACGAATTGGTGGAATGA
>DPOGS206980-PA
MEKYLGKKYKLRSSHNFDEYLKFIEVGLLSRKLVTSLSPISVLTKNEDGSYSLTMITPIRKVVITFQLGVEFSEDRPDGIKVKSTMHLDGDKLIQTQIESNGRKSTHVREFTDKLLTVTTTAEGWDGQCIRIYELVE-