| DPOGS207265 | ||
|---|---|---|
| Transcript | DPOGS207265-TA | 657 bp |
| Protein | DPOGS207265-PA | 218 aa |
| Genomic position | DPSCF300008 - 669429-671256 | |
| RNAseq coverage | 375x (Rank: top 32%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL016335 | 2e-57 | 72.14% | |
| Bombyx | BGIBMGA012129-TA | 5e-60 | 70.88% | |
| Drosophila | Spn-PJ | 7e-20 | 33.33% | |
| EBI UniRef50 | UniRef50_E2B7T8 | 6e-26 | 39.83% | Neurabin-1 n=7 Tax=Formicidae RepID=E2B7T8_HARSA |
| NCBI RefSeq | XP_623895.2 | 3e-29 | 40.25% | PREDICTED: similar to Spinophilin CG16757-PA [Apis mellifera] |
| NCBI nr blastp | gi|340719503 | 6e-28 | 41.67% | PREDICTED: hypothetical protein LOC100646066 [Bombus terrestris] |
| NCBI nr blastx | gi|350410506 | 3e-30 | 41.74% | PREDICTED: hypothetical protein LOC100748486 [Bombus impatiens] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 6.7e-12 | protein binding | |
| KEGG pathway | ||||
| InterPro domain | [130-206] IPR010993 | 6.7e-12 | Sterile alpha motif homology | |
| [129-202] IPR013761 | 1.2e-10 | Sterile alpha motif-type | ||
| [136-198] IPR021129 | 1.9e-06 | Sterile alpha motif, type 1 | ||
| Orthology group | MCL14104 | Multiple-copy universal gene | ||
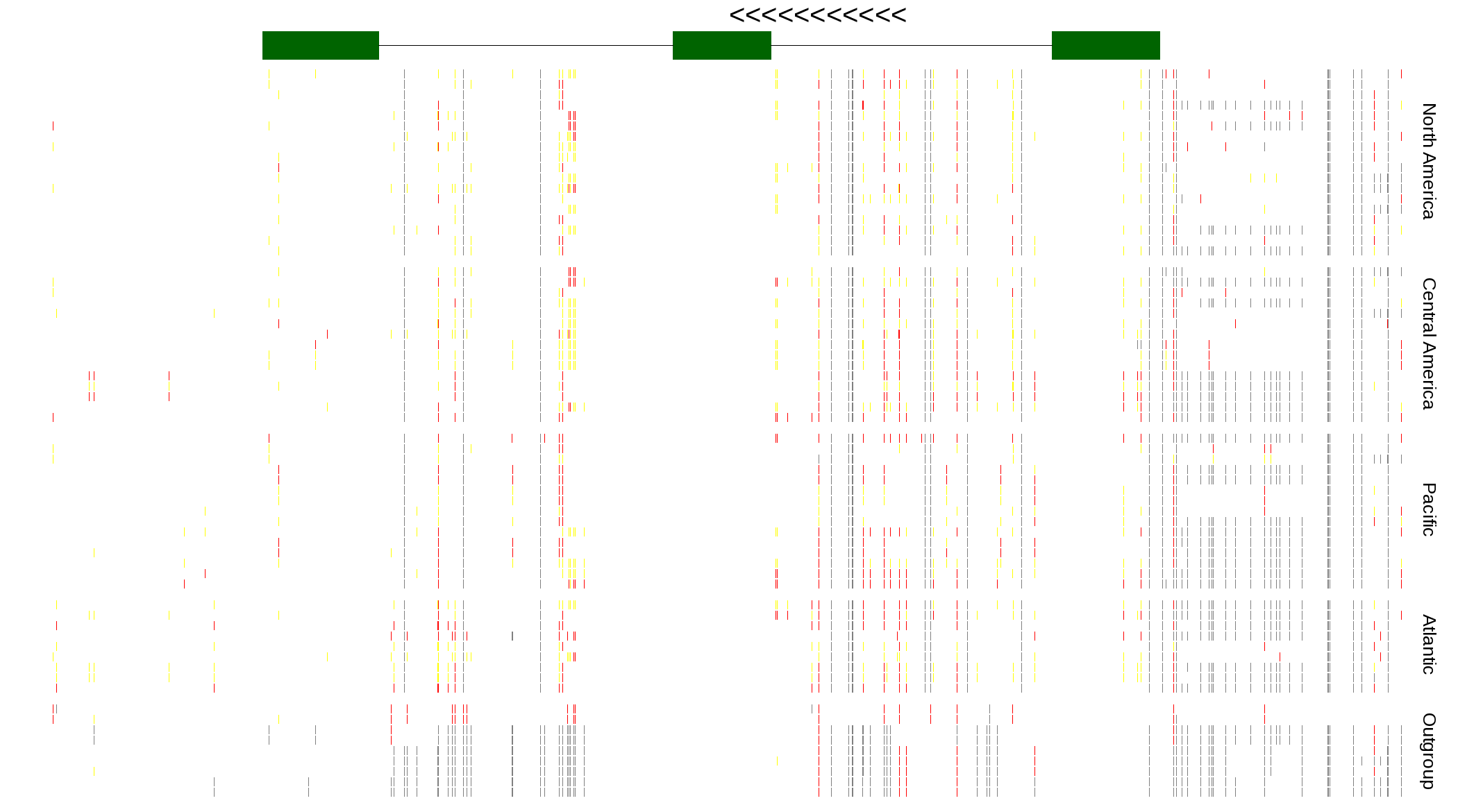
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS207265-TA
ATGCCTCTACATAGAGAGGCAGAAGCCGTATGTTGTTGTGTGAGCGCGGTGGTGGGTGGCCCGGTAGGTCCGCCGGCGTCGCTGGCGGAGCAGCTGAAGCAGGTGTTGGCGGAGCGCGAGCGGCGGCTGGCCAGCGGATCCCCCCACCCTGACCCCGTGCAGCTCACCAACGACCTCGTGGACGAGATACGCCAGGCCGTCAGCGAAGCTAACGCCAGAGTGAAGAAAGCTCCGGCTTGTGTAGTCGGCGGCGAGGTGCCGTGGCAGCCGCTGTCTGCGGGGGCTTCTGAGACCAGCCTGTCGCCGGGTCCCCACGGCCCGCAAGGCCCCCACGGTCCGCAAGGCCCCCACGGACCACACGTTTCACTCACGTCACTAGTCGCCTCGCCCCAGCCAGTCAACCTCTGGACTAAGCAACAGGTGTGGCAGTGGGTATCGAGCCTGGGCGAGGGTCTGGATCGTCACGCGGGTTCGTTTGCGACGCGCGGCGTTGATGGCTCGCTGCTGTTGGCGCTCAGCTCCAGCGACCTGAAGCTGCTGGGTCTGGCGGGGGACGATCGCCGCCGTATGAAAAGACGCCTCAAGGATCTGCGGGCCTCGCACGACAAGTACCTCAAGGCGGTCAAGAAAGCGGAGAAGAAAGCCAAGAAGAAGTAA
>DPOGS207265-PA
MPLHREAEAVCCCVSAVVGGPVGPPASLAEQLKQVLAERERRLASGSPHPDPVQLTNDLVDEIRQAVSEANARVKKAPACVVGGEVPWQPLSAGASETSLSPGPHGPQGPHGPQGPHGPHVSLTSLVASPQPVNLWTKQQVWQWVSSLGEGLDRHAGSFATRGVDGSLLLALSSSDLKLLGLAGDDRRRMKRRLKDLRASHDKYLKAVKKAEKKAKKK-