| DPOGS207772 | ||
|---|---|---|
| Transcript | DPOGS207772-TA | 552 bp |
| Protein | DPOGS207772-PA | 183 aa |
| Genomic position | DPSCF300042 - 187532-188721 | |
| RNAseq coverage | 1926x (Rank: top 6%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL017573 | 2e-99 | 95.63% | |
| Bombyx | BGIBMGA005328-TA | 4e-84 | 90.12% | |
| Drosophila | CG5885-PA | 1e-70 | 66.13% | |
| EBI UniRef50 | UniRef50_Q9VL69 | 1e-68 | 66.13% | CG5885 n=19 Tax=Eumetazoa RepID=Q9VL69_DROME |
| NCBI RefSeq | NP_001040330.1 | 1e-93 | 90.16% | translocon-associated protein gamma isoform 1 [Bombyx mori] |
| NCBI nr blastp | gi|290563285 | 2e-92 | 90.16% | translocon-associated protein gamma isoform 1 [Bombyx mori] |
| NCBI nr blastx | gi|290563285 | 1e-89 | 90.16% | translocon-associated protein gamma isoform 1 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0030176 | 1.2e-122 | integral to endoplasmic reticulum membrane | |
| GO:0006613 | 1.2e-122 | cotranslational protein targeting to membrane | ||
| GO:0005784 | 1.2e-122 | Sec61 translocon complex | ||
| KEGG pathway | nvi:100120928 | 4e-77 | ||
| K13251 (SSR3) | maps-> | Protein processing in endoplasmic reticulum | ||
| InterPro domain | [4-183] IPR009779 | 1.2e-122 | Translocon-associated, gamma subunit | |
| Orthology group | MCL13168 | Single-copy universal gene | ||
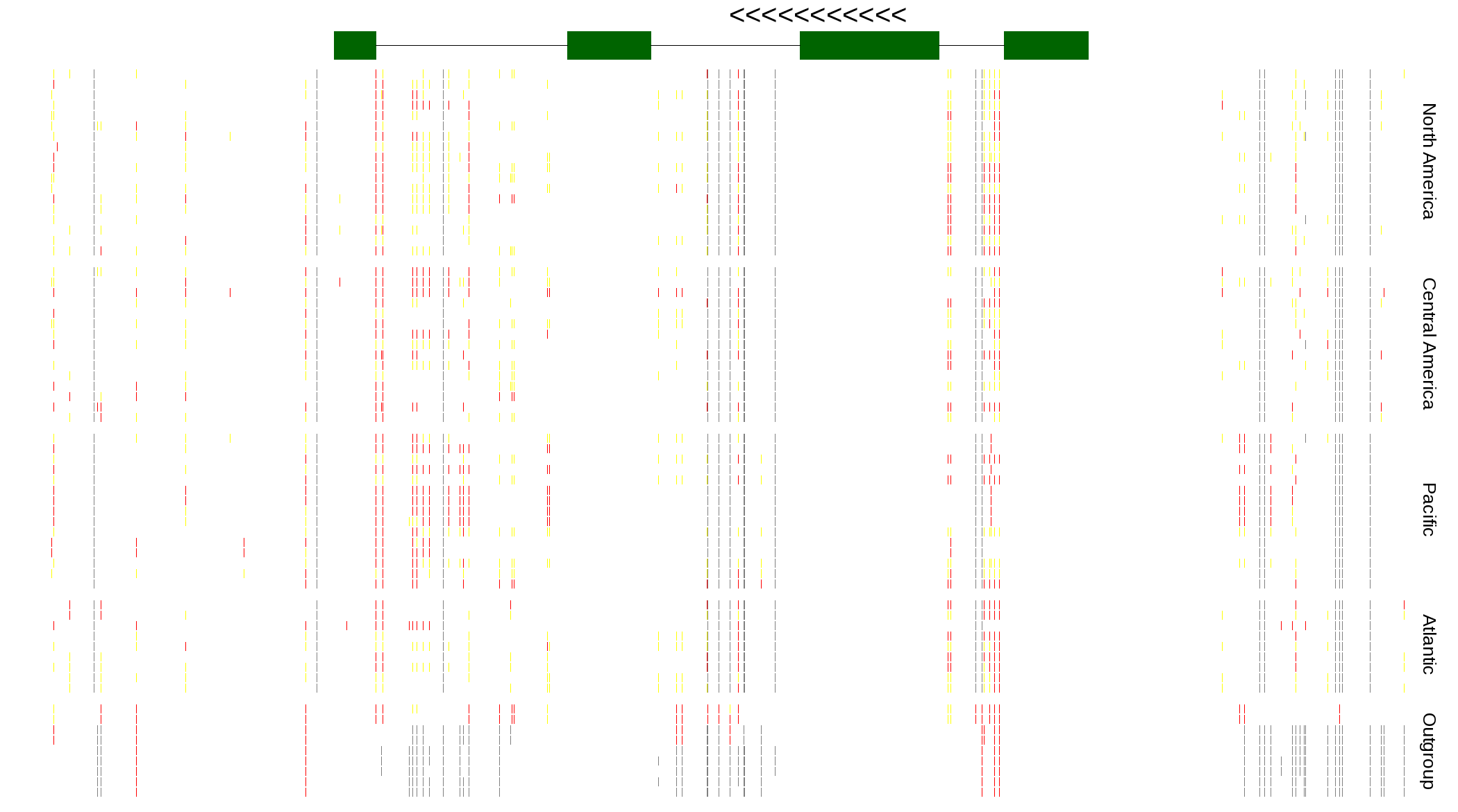
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS207772-TA
ATGTCAGGAAAGGCAAACAAAGCTTTTACGAAGGAGGAAGAGTTGCTGCTACAGGATTTTAGTAGAAATGTATCTACGAAATCTTCTGCCTTATTTTATGGAAATGCATTCATCGTTTCTGCTATACCTATATGGCTGTTCTGGAGAGTTCACGCTTTAGAAGTAAGCTCATCATTGGCTTGGTTTGCTATAGTTACCGTGGCCAGCACTTGGTTGCTTGCTTTAGCTTACCGTAATACCAAGTTCCAATTGAAACACCGTGTCGCTGTACGTCGTGAAGATGCCGTCGCTCGTGAGATGGCTAGAAAGTTGGCTGATGAAAAGAAAATGAGCAGGAAGGAGAAGGATGAAAGAATTTTATGGAAAAAGAATGAAGTTGCTGACTATGAAGCAACAACTTTCTCAATATTTTACAACAATGCTCTATTCTTGACTATTGTTATCCTCGCCAGCTTCTATCTCCTGCGTTCATTCACACCCACCGTTAACTACATTGTATCCCTCACAGTTGCCTCAGGACTTCTGGCTCTTCTGTCTACCGGCTCTAAGTGA
>DPOGS207772-PA
MSGKANKAFTKEEELLLQDFSRNVSTKSSALFYGNAFIVSAIPIWLFWRVHALEVSSSLAWFAIVTVASTWLLALAYRNTKFQLKHRVAVRREDAVAREMARKLADEKKMSRKEKDERILWKKNEVADYEATTFSIFYNNALFLTIVILASFYLLRSFTPTVNYIVSLTVASGLLALLSTGSK-