| DPOGS208043 | ||
|---|---|---|
| Transcript | DPOGS208043-TA | 444 bp |
| Protein | DPOGS208043-PA | 147 aa |
| Genomic position | DPSCF300203 + 164017-165414 | |
| RNAseq coverage | 22170x (Rank: top 0%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL014605 | 2e-73 | 97.96% | |
| Bombyx | BGIBMGA001471-TA | 1e-81 | 98.64% | |
| Drosophila | RpS15-PA | 6e-68 | 87.59% | |
| EBI UniRef50 | UniRef50_F0UE73 | 1e-64 | 76.71% | 40S ribosomal protein S15 n=2 Tax=Ajellomyces capsulatus RepID=F0UE73_AJEC8 |
| NCBI RefSeq | NP_001037209.1 | 4e-80 | 98.64% | ribosomal protein S15 [Bombyx mori] |
| NCBI nr blastp | gi|15213818 | 1e-79 | 100.00% | ribosomal protein S15 [Spodoptera frugiperda] |
| NCBI nr blastx | gi|15213818 | 7e-79 | 100.00% | ribosomal protein S15 [Spodoptera frugiperda] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005840 | 1.8e-105 | ribosome | |
| GO:0006412 | 1.8e-105 | translation | ||
| GO:0003735 | 1.8e-105 | structural constituent of ribosome | ||
| GO:0015935 | 1.3e-68 | small ribosomal subunit | ||
| KEGG pathway | nvi:100113507 | 2e-78 | ||
| K02958 (RP-S15e, RPS15) | maps-> | Ribosome | ||
| InterPro domain | [2-148] IPR002222 | 1.8e-105 | Ribosomal protein S19/S15 | |
| [14-147] IPR005713 | 1.3e-68 | Ribosomal protein S19A/S15e | ||
| [48-134] IPR023575 | 6.7e-37 | Ribosomal protein S19, superfamily | ||
| Orthology group | MCL14498 | Single-copy universal gene | ||
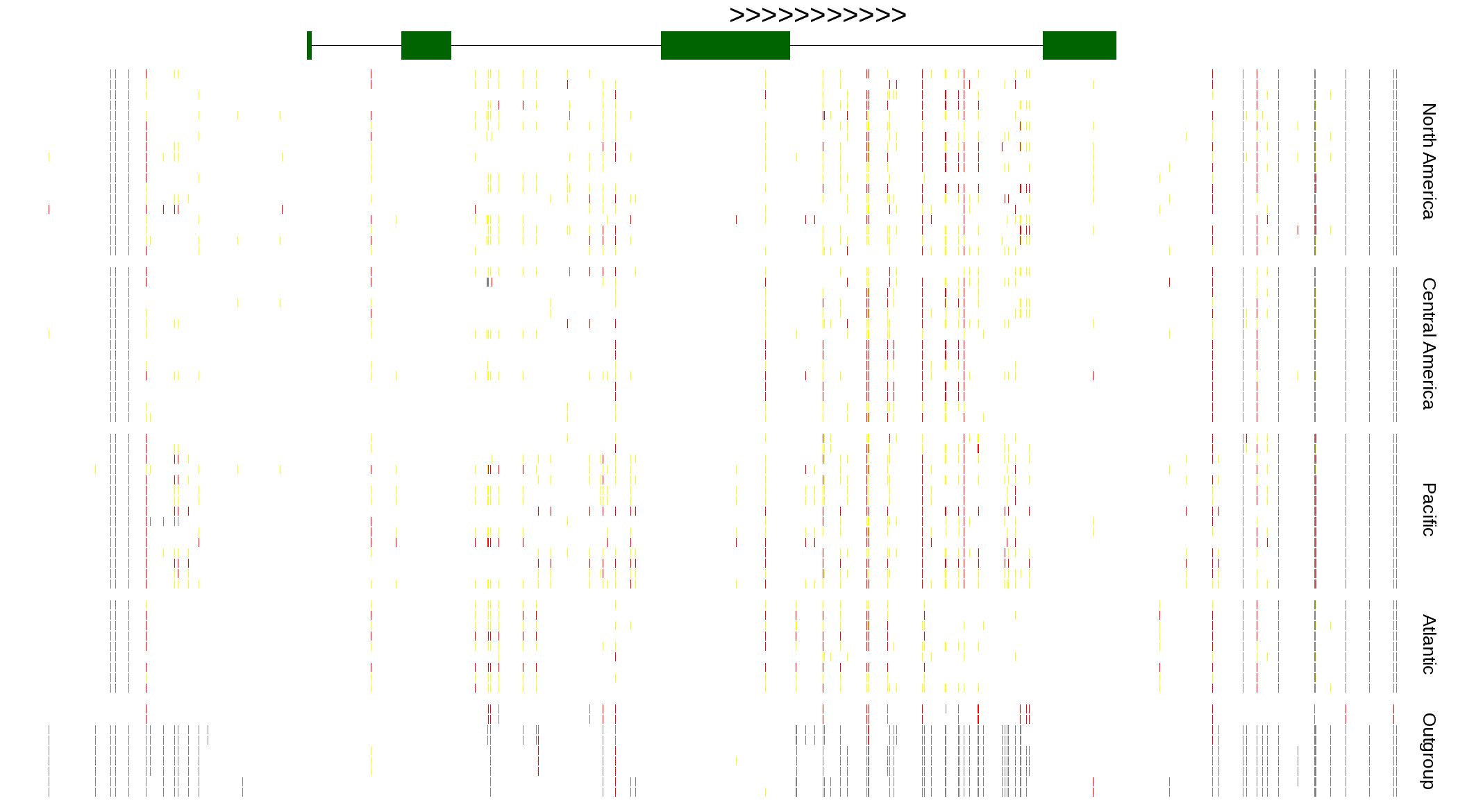
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS208043-TA
ATGGCTGAGGTCGACGAAACTCTTAAAAAGAAGCGTACCTTCAGGAAGTTTACCTTCCGAGGTGTCGACTTGGACCAGCTTCTTGATATGCCAAATGAACAGCTCATGGAGTTGATGCACGCTCGCGCCCGTAGGCGTTTCGCCCGCGGTCTCAAACGCAAGCCCATGGCGCTGGTGAAGAAACTGAGACGCGCCAAGAAGGAAGCTCCTCCAAATGAGAAGCCCGAGATTGTCAAGACACACTTAAGAAACATGATCATTGTGCCAGAGATGGTCGGATCCATTGTTGGTATTTACAACGGCAAGACTTTCAATCAGGTTGAAATCAAGCCTGAGATGATTGGTCACTACTTGGGTGAATTCTCTGTGACATACAAACCCGTAAAGCACGGTCGGCCGGGTATTGGTGCCACCCACAGTTCCAGGTTCATCCCTCTGAAGTAG
>DPOGS208043-PA
MAEVDETLKKKRTFRKFTFRGVDLDQLLDMPNEQLMELMHARARRRFARGLKRKPMALVKKLRRAKKEAPPNEKPEIVKTHLRNMIIVPEMVGSIVGIYNGKTFNQVEIKPEMIGHYLGEFSVTYKPVKHGRPGIGATHSSRFIPLK-