| DPOGS208119 | ||
|---|---|---|
| Transcript | DPOGS208119-TA | 555 bp |
| Protein | DPOGS208119-PA | 184 aa |
| Genomic position | DPSCF300154 - 91570-92636 | |
| RNAseq coverage | 68x (Rank: top 67%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012271 | 6e-105 | 97.83% | |
| Bombyx | BGIBMGA006771-TA | 2e-104 | 97.83% | |
| Drosophila | CG17266-PA | 2e-93 | 86.41% | |
| EBI UniRef50 | UniRef50_C4QM49 | 1e-79 | 73.96% | Peptidyl-prolyl cis-trans isomerase h, ppih,putative n=9 Tax=Eukaryota RepID=C4QM49_SCHMA |
| NCBI RefSeq | XP_001120773.1 | 1e-97 | 91.85% | PREDICTED: similar to CG17266-PA [Apis mellifera] |
| NCBI nr blastp | gi|383848460 | 1e-96 | 92.39% | PREDICTED: peptidyl-prolyl cis-trans isomerase H-like [Megachile rotundata] |
| NCBI nr blastx | gi|383848460 | 4e-99 | 92.39% | PREDICTED: peptidyl-prolyl cis-trans isomerase H-like [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006457 | 2.3e-72 | protein folding | |
| GO:0003755 | 2.3e-72 | peptidyl-prolyl cis-trans isomerase activity | ||
| KEGG pathway | ame:726409 | 4e-97 | ||
| K09567 (PPIH, CYPH) | maps-> | Spliceosome | ||
| InterPro domain | [11-183] IPR002130 | 2.3e-72 | Peptidyl-prolyl cis-trans isomerase, cyclophilin-type | |
| [1-184] IPR015891 | 3.6e-70 | Cyclophilin-like | ||
| Orthology group | MCL12953 | Single-copy universal gene | ||
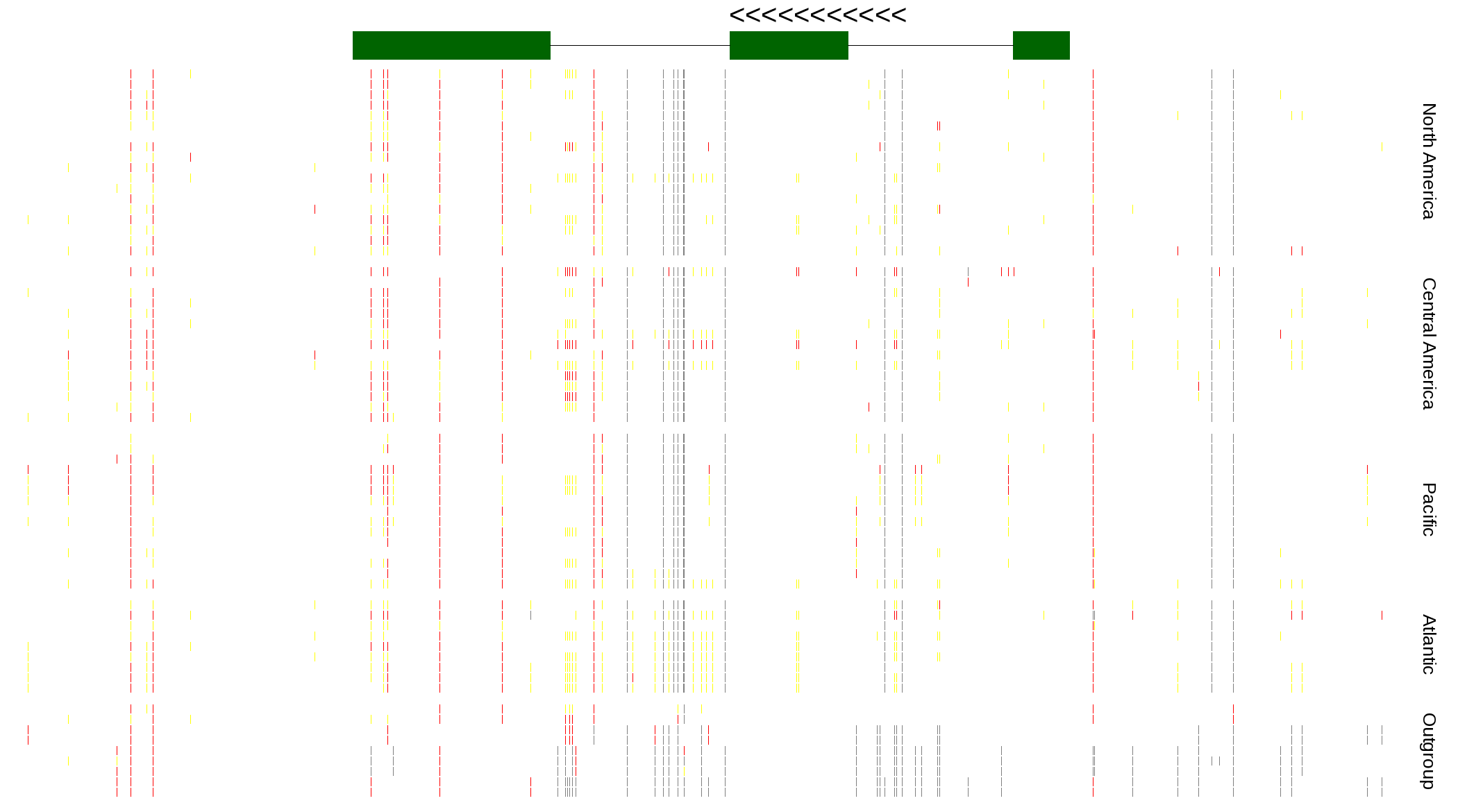
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS208119-TA
ATGCCGACATGGAATCAAATACAAACACAACTCAGAAATCCAAATAATCCAGTTGTATTCTTTGATATTACAGTCGGCACAACTGAAATTGGTCGCATGATATTTGAACTTTTTGCTGATGTTGTACCAAAGACAAGTGAAAACTTCAGAGAATTCTGCACAGGTGAATACAGAAGAGATGGTGTTCCACTTGGTTTCAAAGGCGCAACTTTTCATAGAGTAATAAAAGACTTTATGATACAAGGGGGAGATTTTGTAAATGGTGATGGCACAGGTGTGATGAGTATTTACGGAGGTAGTACATTTGCTGATGAGAACTTTGTAGTAAAGCATGATTCCCCAGGATTATTATCAATGGCAAATAGTGGCAAGGACACAAATGGCTGTCAGTTTTTCATAACTTGCGCCAAGTGTAATTTCTTAGATAATAAGCATGTTGTGTTCGGAAGAGTTATAGATGGTTTATTAGTTATGAGAAAAATTGAAAATGTACCCACAGGACCAAACAATAAACCCAAAATACCAGTGACAATATCACAGTGTGGACAAATGTAA
>DPOGS208119-PA
MPTWNQIQTQLRNPNNPVVFFDITVGTTEIGRMIFELFADVVPKTSENFREFCTGEYRRDGVPLGFKGATFHRVIKDFMIQGGDFVNGDGTGVMSIYGGSTFADENFVVKHDSPGLLSMANSGKDTNGCQFFITCAKCNFLDNKHVVFGRVIDGLLVMRKIENVPTGPNNKPKIPVTISQCGQM-