| DPOGS208284 | ||
|---|---|---|
| Transcript | DPOGS208284-TA | 570 bp |
| Protein | DPOGS208284-PA | 189 aa |
| Genomic position | DPSCF300079 + 173845-175146 | |
| RNAseq coverage | 37x (Rank: top 73%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL002140 | 5e-43 | 63.64% | |
| Bombyx | BGIBMGA007914-TA | 1e-18 | 52.70% | |
| Drosophila | Hr39-PB | 2e-20 | 52.70% | |
| EBI UniRef50 | UniRef50_B3MMC7 | 1e-18 | 52.70% | GF14318 n=1 Tax=Drosophila ananassae RepID=B3MMC7_DROAN |
| NCBI RefSeq | XP_002057789.1 | 3e-19 | 52.70% | GJ18326 [Drosophila virilis] |
| NCBI nr blastp | gi|195398359 | 4e-18 | 52.70% | GJ18326 [Drosophila virilis] |
| NCBI nr blastx | gi|195443384 | 2e-16 | 54.17% | GK18731 [Drosophila willistoni] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005634 | 6.4e-28 | nucleus | |
| GO:0006355 | 6.4e-28 | regulation of transcription, DNA-dependent | ||
| GO:0043565 | 6.4e-28 | sequence-specific DNA binding | ||
| GO:0008270 | 6.4e-28 | zinc ion binding | ||
| GO:0003700 | 6.4e-28 | sequence-specific DNA binding transcription factor activity | ||
| KEGG pathway | ||||
| InterPro domain | [32-108] IPR001628 | 6.4e-28 | Zinc finger, nuclear hormone receptor-type | |
| [33-111] IPR013088 | 5.4e-23 | Zinc finger, NHR/GATA-type | ||
| [124-182] IPR007588 | 7.2e-08 | Zinc finger, FLYWCH-type | ||
| Orthology group | MCL34741 | Lepidoptera specific | ||
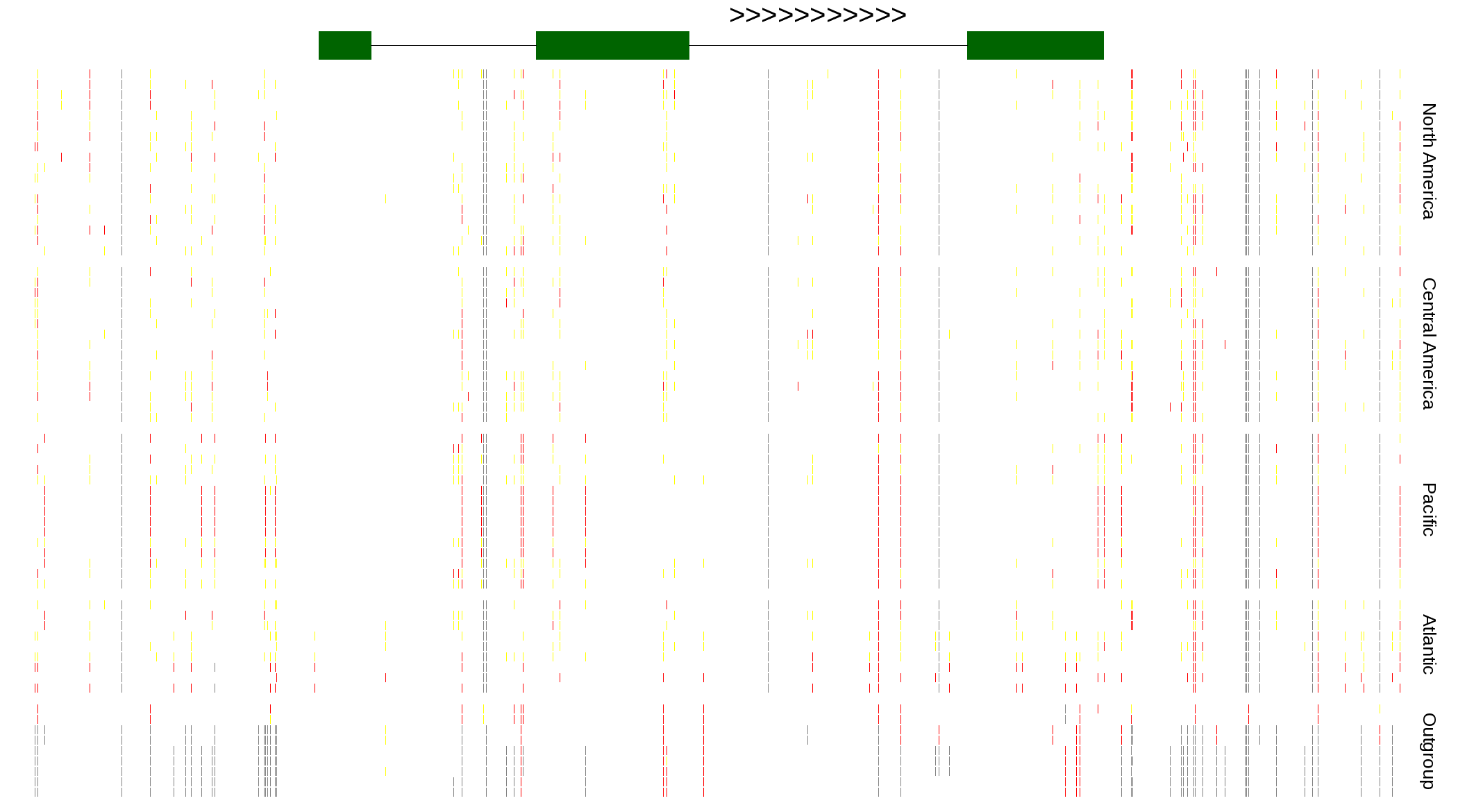
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS208284-TA
ATGGTTGTCGGTGGCTATAGATACTCGCTGCATTACTCCAGGAATAATAGAGAGAGGTGGATGTGCAGCAGACGGAGTTACTACGGCTGGCCGATAACATCTTGTCCGATCTGCGGCGATGCGGTGAATGGGATACATTACGGTATATTCACGTGTGAGTCGTGTAAATCTTTCTTCAGGCGTACCTTACAGACGAACAAGCAAGCGGGTTACGTGTGCAGATATAAGAGACGGAAGCGACCTTGTGATATGACCGTACAAACCAGATCTTTCTGCAGATTCTGCAGGTTCAATAGATGCTTGGAAGCCGGGATGCAACCCGGTCTGGTCAAGATGATTAAAGATTTCGTGGTACCGAGGTACAGCTGGTCGCGAGCCGGTAATGCTATCATCATTCTGGGAGACTACAGGTTCAAAAAGCGATCTAAGCACATCAATCAGAAACAGGAGACGCATTGGGTGTGTAACAAATGTGATAAGGGATGTAAAGCGAAGCTGACAACTCTCAATGACGCCATCATCAAGATGTACAACGTCCACAAACATGATGGTTCCAGAAAATTAAAGTGA
>DPOGS208284-PA
MVVGGYRYSLHYSRNNRERWMCSRRSYYGWPITSCPICGDAVNGIHYGIFTCESCKSFFRRTLQTNKQAGYVCRYKRRKRPCDMTVQTRSFCRFCRFNRCLEAGMQPGLVKMIKDFVVPRYSWSRAGNAIIILGDYRFKKRSKHINQKQETHWVCNKCDKGCKAKLTTLNDAIIKMYNVHKHDGSRKLK-