| DPOGS208306 | ||
|---|---|---|
| Transcript | DPOGS208306-TA | 420 bp |
| Protein | DPOGS208306-PA | 139 aa |
| Genomic position | DPSCF300948 - 3122-5154 | |
| RNAseq coverage | 55x (Rank: top 69%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL020310 | 2e-15 | 33.94% | |
| Bombyx | BGIBMGA003228-TA | 5e-18 | 82.22% | |
| Drosophila | Ance-3-PB | 2e-17 | 41.51% | |
| EBI UniRef50 | UniRef50_UPI0002063A41 | 2e-31 | 54.13% | UPI0002063A41 related cluster n=1 Tax=unknown RepID=UPI0002063A41 |
| NCBI RefSeq | XP_001602098.1 | 3e-32 | 47.76% | PREDICTED: similar to ENSANGP00000007115 [Nasonia vitripennis] |
| NCBI nr blastp | gi|328779592 | 8e-31 | 54.13% | PREDICTED: angiotensin-converting enzyme-like [Apis mellifera] |
| NCBI nr blastx | gi|91087059 | 2e-29 | 41.73% | PREDICTED: similar to angiotensin-converting enzyme 8 (AGAP007982-PA) [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016020 | 8.1e-35 | membrane | |
| GO:0008241 | 8.1e-35 | peptidyl-dipeptidase activity | ||
| GO:0008237 | 8.1e-35 | metallopeptidase activity | ||
| GO:0006508 | 8.1e-35 | proteolysis | ||
| KEGG pathway | nvi:100118011 | 9e-32 | ||
| K01283 (E3.4.15.1, ACE) | maps-> | Chagas disease | ||
| Renin-angiotensin system | ||||
| Hypertrophic cardiomyopathy (HCM) | ||||
| InterPro domain | [4-105] IPR001548 | 8.1e-35 | Peptidase M2, peptidyl-dipeptidase A | |
| Orthology group | ||||
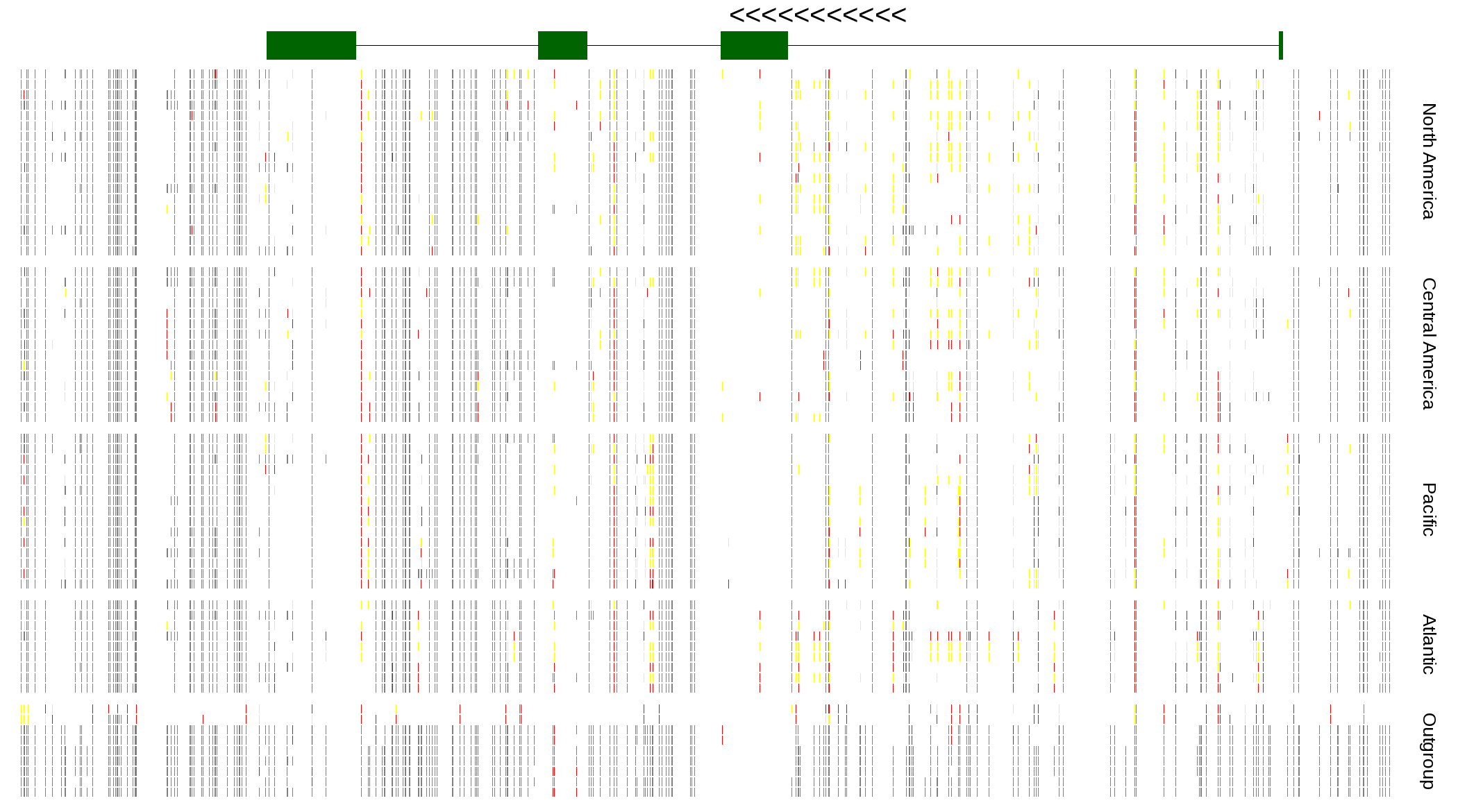
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS208306-TA
ATGTATAAAGATTACATAAAATATTTCCTAGCTACGGTCCTCGAGTTCCAGGTCTTCGAGCAGCTGTGCGCTGCTGCAGGTCACAATGGACACCTGCACGAGTGTGACATCTACAGGAGCAGAGACGCTGGCAGACTTCTGGGTGAGACGATGCAAATAGGAGCGTCAAAACCAGCGTCGGACGTCATAAGAATAATGACTAGAGGAAAGACGAACAGAATATCGCCGGAATCTATTGTGAAATTCTTCCGACCCCTGGAACTCTGGCTGAGAGTTCAGAATCGTGATGAGGAGGTGATAGGATGGAACTCAAACTACGAGGACGTTGCTCTCTTCGCTCCGCAGAGGGCGCTGGCTAGTGACAGTCATCTGTCACCTGTTGTAACACTCGCTTCATTCATTGTTTTATTTTATAAATAA
>DPOGS208306-PA
MYKDYIKYFLATVLEFQVFEQLCAAAGHNGHLHECDIYRSRDAGRLLGETMQIGASKPASDVIRIMTRGKTNRISPESIVKFFRPLELWLRVQNRDEEVIGWNSNYEDVALFAPQRALASDSHLSPVVTLASFIVLFYK-