| DPOGS210794 | ||
|---|---|---|
| Transcript | DPOGS210794-TA | 681 bp |
| Protein | DPOGS210794-PA | 226 aa |
| Genomic position | DPSCF300027 - 1164537-1168196 | |
| RNAseq coverage | 1145x (Rank: top 11%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL007740 | 4e-30 | 70.00% | |
| Bombyx | BGIBMGA007109-TA | 2e-107 | 86.92% | |
| Drosophila | synaptogyrin-PA | 5e-54 | 47.49% | |
| EBI UniRef50 | UniRef50_F4W8F8 | 9e-70 | 56.96% | Synaptogyrin-2 n=4 Tax=Endopterygota RepID=F4W8F8_ACREC |
| NCBI RefSeq | XP_001605760.1 | 1e-76 | 67.61% | PREDICTED: similar to synaptogyrin, putative [Nasonia vitripennis] |
| NCBI nr blastp | gi|307199468 | 2e-75 | 63.72% | Synaptogyrin-2 [Harpegnathos saltator] |
| NCBI nr blastx | gi|345485430 | 1e-84 | 67.54% | PREDICTED: synaptogyrin-2-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | tgu:100223478 | 1e-34 | ||
| K04403 (MAP3K7IP1, TAB1) | maps-> | Leishmaniasis | ||
| Toll-like receptor signaling pathway | ||||
| MAPK signaling pathway | ||||
| NOD-like receptor signaling pathway | ||||
| InterPro domain | [24-165] IPR021128 | 1.3e-19 | MARVEL-like domain | |
| Orthology group | MCL10835 | Multiple-copy universal gene | ||
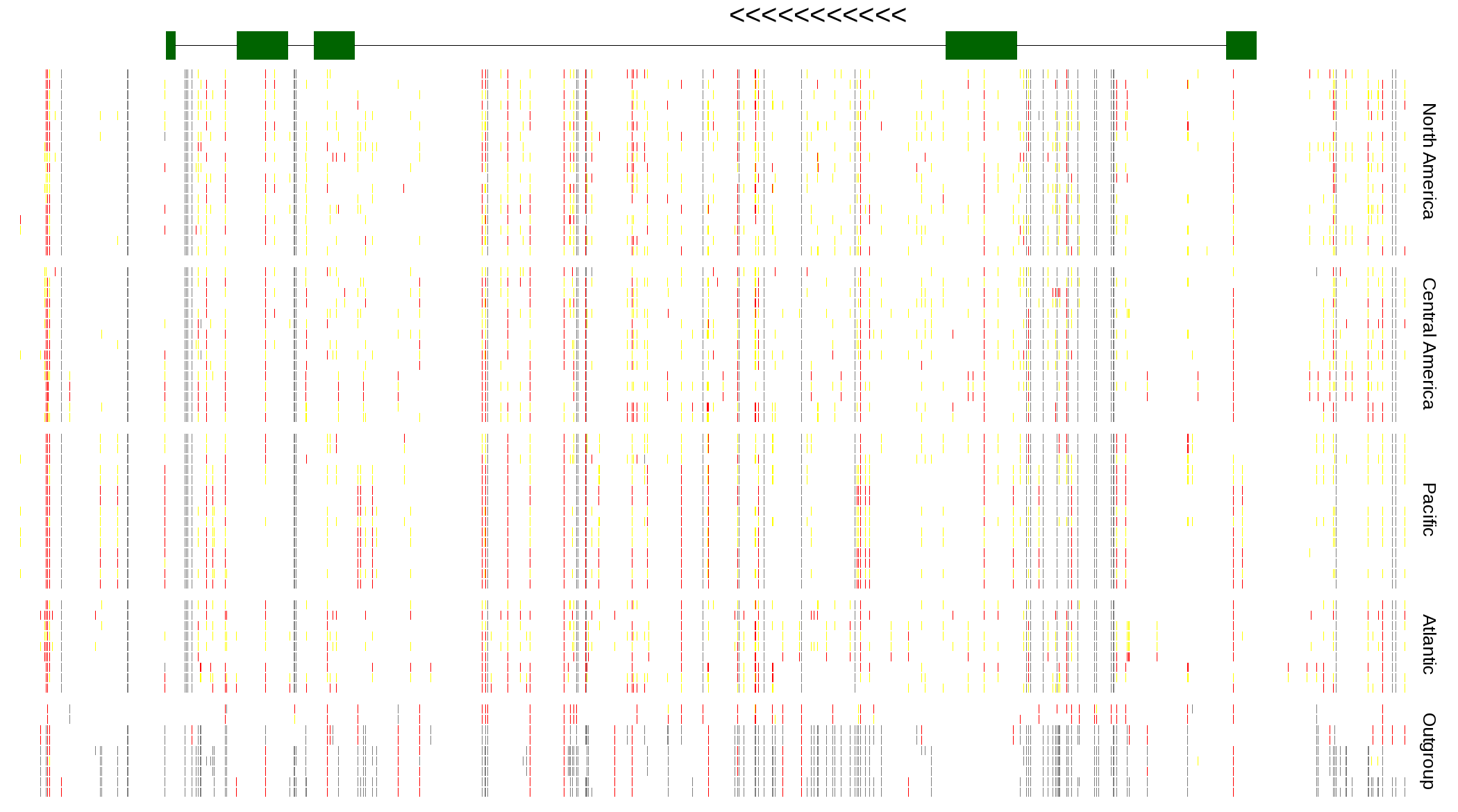
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS210794-TA
ATGGAGTCAGCTGGTGCTTATGGTGGTGGCAAAGCTGGTGGAGCCTTCGATCCTCAGGCTTTTGTTCAAAAGCCTCCGGTTATTGTTCGCTCCGTTTGTTGGCTGTTCAGTGTGATTGTAATGGGCTGTATATCTGCCAAGGGCTGGTACATCAGCAAAACAGACGGCAAAGAGTACTGTGTCTACAACGATGATACAAATGCTTGCAATTATGGAGTCGGAATCTCAGTCATAGCATTTGTTGCATCTATAGCTTTCATTGTGGGGGAGTACTTGTTTGAACAAATGTCCTCCGTGAAGACAAGAAAACATTATGTTCTTGCTGATATGGGATTTTCTGCGTTCTGGGCGTTTTTATATTTCGTGGGCTTCTGTTATTTGTCCAACGCCTGGGGTAAGACTGAGAACCCGCCGATGGGTACAGCCAACAATATGCAGGGAGCCATCGCGTTCTGTTTCTTCTCGATTTTTGCTTGGGTAGCGTGTGCGTTCTTCGCATTCCAACGCTTCCGTATGGGCGCTGACGCGGCCTTCGCTCCCGCATACGAAGTTGAAGGCGCTGGCAGCCCGGGTACCTTCCCCGCCTACCCCGGCGCTCCGGACCCACAGCCGGCGTACAGCGACCCACCCTTCCAACAACAACACACTGGTAACATGGACTACACCGCCCCTACATACTGA
>DPOGS210794-PA
MESAGAYGGGKAGGAFDPQAFVQKPPVIVRSVCWLFSVIVMGCISAKGWYISKTDGKEYCVYNDDTNACNYGVGISVIAFVASIAFIVGEYLFEQMSSVKTRKHYVLADMGFSAFWAFLYFVGFCYLSNAWGKTENPPMGTANNMQGAIAFCFFSIFAWVACAFFAFQRFRMGADAAFAPAYEVEGAGSPGTFPAYPGAPDPQPAYSDPPFQQQHTGNMDYTAPTY-