| DPOGS210995 | ||
|---|---|---|
| Transcript | DPOGS210995-TA | 288 bp |
| Protein | DPOGS210995-PA | 95 aa |
| Genomic position | DPSCF300004 + 276906-277989 | |
| RNAseq coverage | 3x (Rank: top 90%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL025027 | 2e-34 | 87.32% | |
| Bombyx | BGIBMGA006476-TA | 5e-31 | 80.28% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_E2B7P0 | 2e-12 | 52.54% | Protein CREG1 n=2 Tax=Harpegnathos saltator RepID=E2B7P0_HARSA |
| NCBI RefSeq | XP_001600388.1 | 2e-13 | 59.32% | PREDICTED: similar to conserved hypothetical protein [Nasonia vitripennis] |
| NCBI nr blastp | gi|307212603 | 7e-12 | 52.54% | Protein CREG1 [Harpegnathos saltator] |
| NCBI nr blastx | gi|345491960 | 1e-12 | 59.32% | PREDICTED: protein CREG1-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| InterPro domain | [24-87] IPR009002 | 5e-13 | FMN-binding split barrel-related | |
| Orthology group | ||||
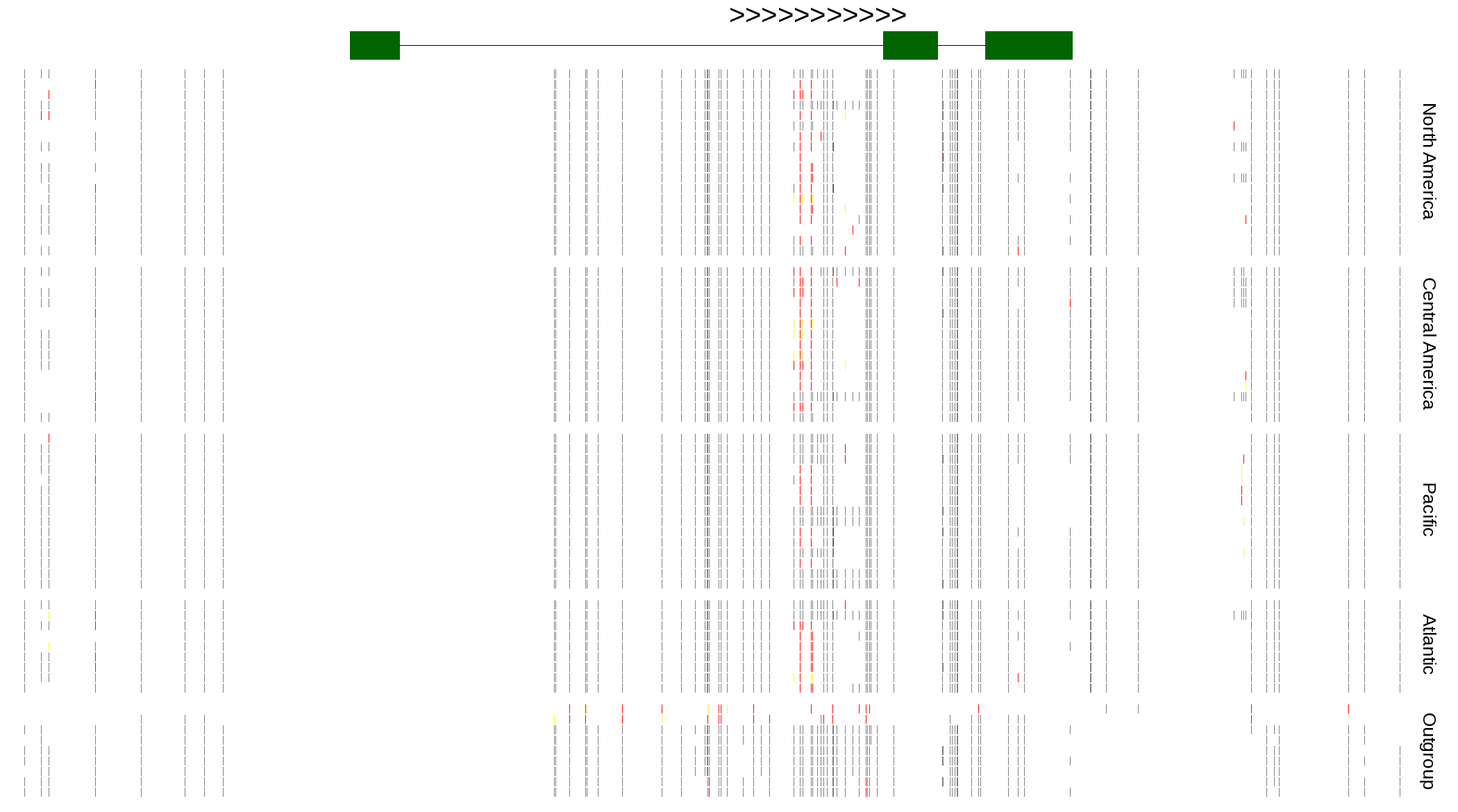
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS210995-TA
ATGTTATTATTCCCCAAACATACGCTACTATCACATGAGTTAATGTCCTTTAGTAAATTGCAAGAGTTTGCTAAAGTTAAGGAAAACACACCAGAATACACATTCGCAAAGGCTGCATTATTCGAAAGGCACCCGGACATGGCGAACTTTCCGCCTGATCACAATTGGTTTGTGGCTAAAATGAAGATAGCCCAAATAGCTATGGTCGATTGGTTCGGTGGTGCTAAATATGTTCCTGTTAAAGATTACCTCGCCTACAATTATACTGGGATGGAGATGTTGCATTAA
>DPOGS210995-PA
MLLFPKHTLLSHELMSFSKLQEFAKVKENTPEYTFAKAALFERHPDMANFPPDHNWFVAKMKIAQIAMVDWFGGAKYVPVKDYLAYNYTGMEMLH-