| DPOGS211301 | ||
|---|---|---|
| Transcript | DPOGS211301-TA | 636 bp |
| Protein | DPOGS211301-PA | 211 aa |
| Genomic position | DPSCF300125 - 268496-277661 | |
| RNAseq coverage | 2105x (Rank: top 6%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL016476 | 5e-118 | 98.56% | |
| Bombyx | BGIBMGA005176-TA | 6e-91 | 99.38% | |
| Drosophila | Snap25-PA | 5e-99 | 83.18% | |
| EBI UniRef50 | UniRef50_D6WT89 | 3e-103 | 85.31% | Synaptosomal-associated protein n=7 Tax=Coelomata RepID=D6WT89_TRICA |
| NCBI RefSeq | XP_974721.1 | 2e-107 | 88.15% | PREDICTED: similar to AGAP001394-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|91087735 | 3e-106 | 88.15% | PREDICTED: similar to AGAP001394-PA [Tribolium castaneum] |
| NCBI nr blastx | gi|91087735 | 1e-106 | 88.15% | PREDICTED: similar to AGAP001394-PA [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 8.5e-16 | protein binding | |
| KEGG pathway | tca:663588 | 5e-107 | ||
| K08508 (SNAP23) | maps-> | SNARE interactions in vesicular transport | ||
| InterPro domain | [97-148] IPR000928 | 6.8e-18 | SNAP-25 | |
| [152-210] IPR000727 | 8.5e-16 | Target SNARE coiled-coil domain | ||
| Orthology group | MCL12458 | Single-copy universal gene | ||
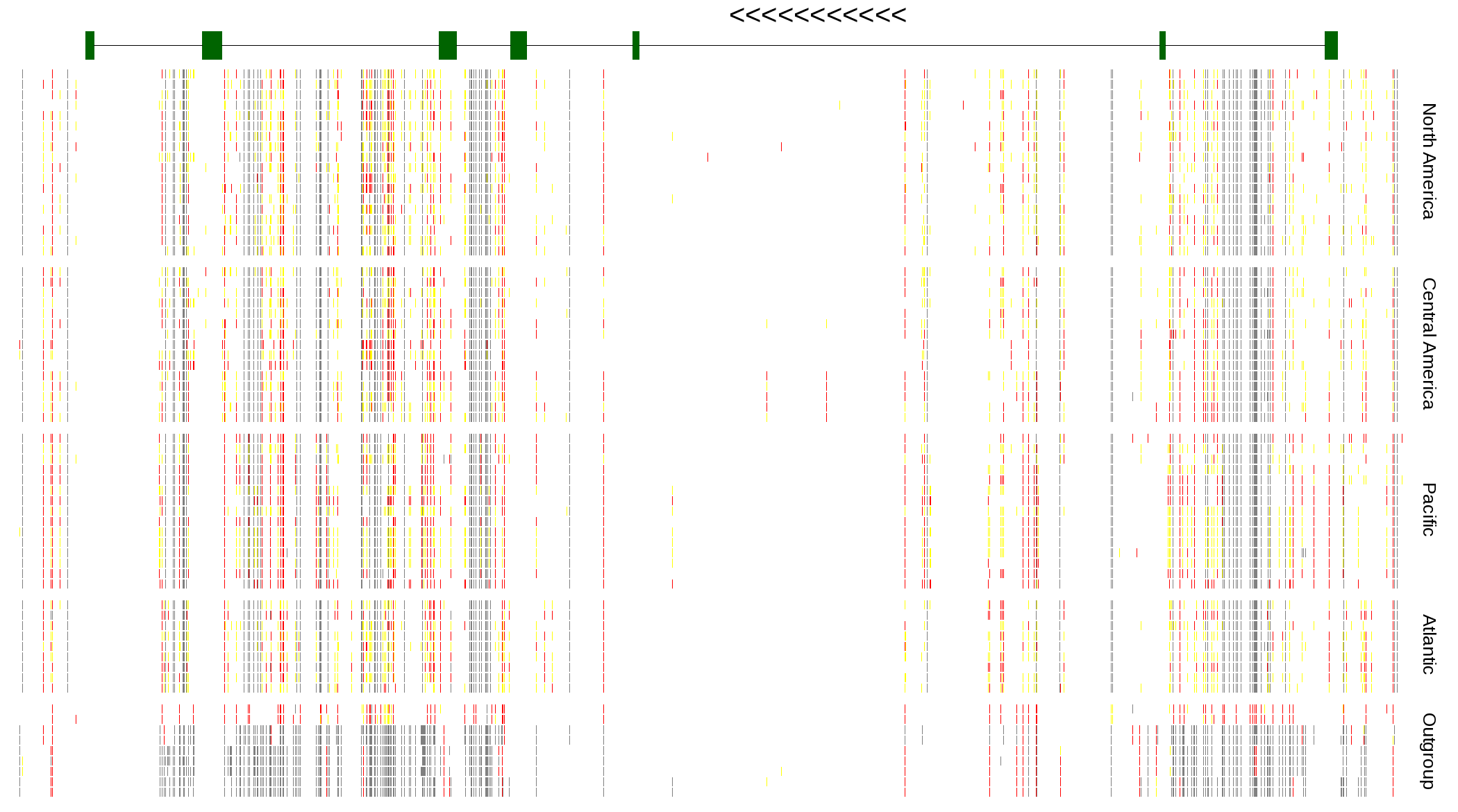
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS211301-TA
ATGCCTTCAGCAGCGCCCCCAGCAGAAAATGGCGCGCCGCGCAGCGAACTTGAGCAATTGCAATTACGAGCCGGCCAGGTCACGGATGAATCCCTGGAGTCGACCCGGCGTATGATGCAACTATGCGAAGAGAGTCACGAGGTGGGCATGAAAACCCTGGTCATGCTGGATGAACAGGGGGAACAACTGGACAGAATCGAGGAGGGTATGGATCAAATTAACGCTGACATGCGTGAGGCTGAGAAGAATCTCTCGGGGATGGAGAAATGTTGTGGCATTTGCGTTCTACCTTGCAACAAGGGGGCGTCATTCAAGGAAGATGATGGTACATGGAAAGGGAACGATGATGGTAAGGTGGTGAACAACCAGCCACAGCGTGTTATGGACGAGCGGAACGGCATCGGACCACAAGCTGGATATATTGGCAGGATAACAAACGACGCTCGCGAAGACGAGATGGAAGAGAACATGGGCCAGGTGAACACCATGATCGGTAACTTACGTAACATGGCCATTGACATGGGCTCCGAGCTGGAGAACCAGAATAGACAGATCGATCGTATCAACCGCAAGGGCGAATCGAACGAAACGAGGATAACGCTCGCCAACCAGCGCGCACACGAGCTCCTCAAGTAA
>DPOGS211301-PA
MPSAAPPAENGAPRSELEQLQLRAGQVTDESLESTRRMMQLCEESHEVGMKTLVMLDEQGEQLDRIEEGMDQINADMREAEKNLSGMEKCCGICVLPCNKGASFKEDDGTWKGNDDGKVVNNQPQRVMDERNGIGPQAGYIGRITNDAREDEMEENMGQVNTMIGNLRNMAIDMGSELENQNRQIDRINRKGESNETRITLANQRAHELLK-