| DPOGS211437 | ||
|---|---|---|
| Transcript | DPOGS211437-TA | 588 bp |
| Protein | DPOGS211437-PA | 195 aa |
| Genomic position | DPSCF300223 - 267967-269674 | |
| RNAseq coverage | 8148x (Rank: top 2%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL015335 | 3e-91 | 82.05% | |
| Bombyx | BGIBMGA002186-TA | 2e-107 | 90.77% | |
| Drosophila | Jafrac1-PA | 4e-88 | 78.53% | |
| EBI UniRef50 | UniRef50_P32119 | 2e-79 | 70.31% | Peroxiredoxin-2 n=216 Tax=root RepID=PRDX2_HUMAN |
| NCBI RefSeq | NP_001037083.1 | 5e-106 | 90.77% | thiol peroxiredoxin [Bombyx mori] |
| NCBI nr blastp | gi|112982996 | 7e-105 | 90.77% | thiol peroxiredoxin [Bombyx mori] |
| NCBI nr blastx | gi|112982996 | 2e-103 | 90.77% | thiol peroxiredoxin [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016209 | 1.2e-39 | antioxidant activity | |
| GO:0055114 | 1.2e-39 | oxidation-reduction process | ||
| GO:0016491 | 1.2e-39 | oxidoreductase activity | ||
| GO:0051920 | 1.9e-15 | peroxiredoxin activity | ||
| KEGG pathway | ||||
| InterPro domain | [4-191] IPR012336 | 4.2e-81 | Thioredoxin-like fold | |
| [7-137] IPR000866 | 1.2e-39 | Alkyl hydroperoxide reductase subunit C/ Thiol specific antioxidant | ||
| [158-191] IPR019479 | 1.9e-15 | Peroxiredoxin, C-terminal | ||
| Orthology group | MCL11140 | Multiple-copy universal gene | ||
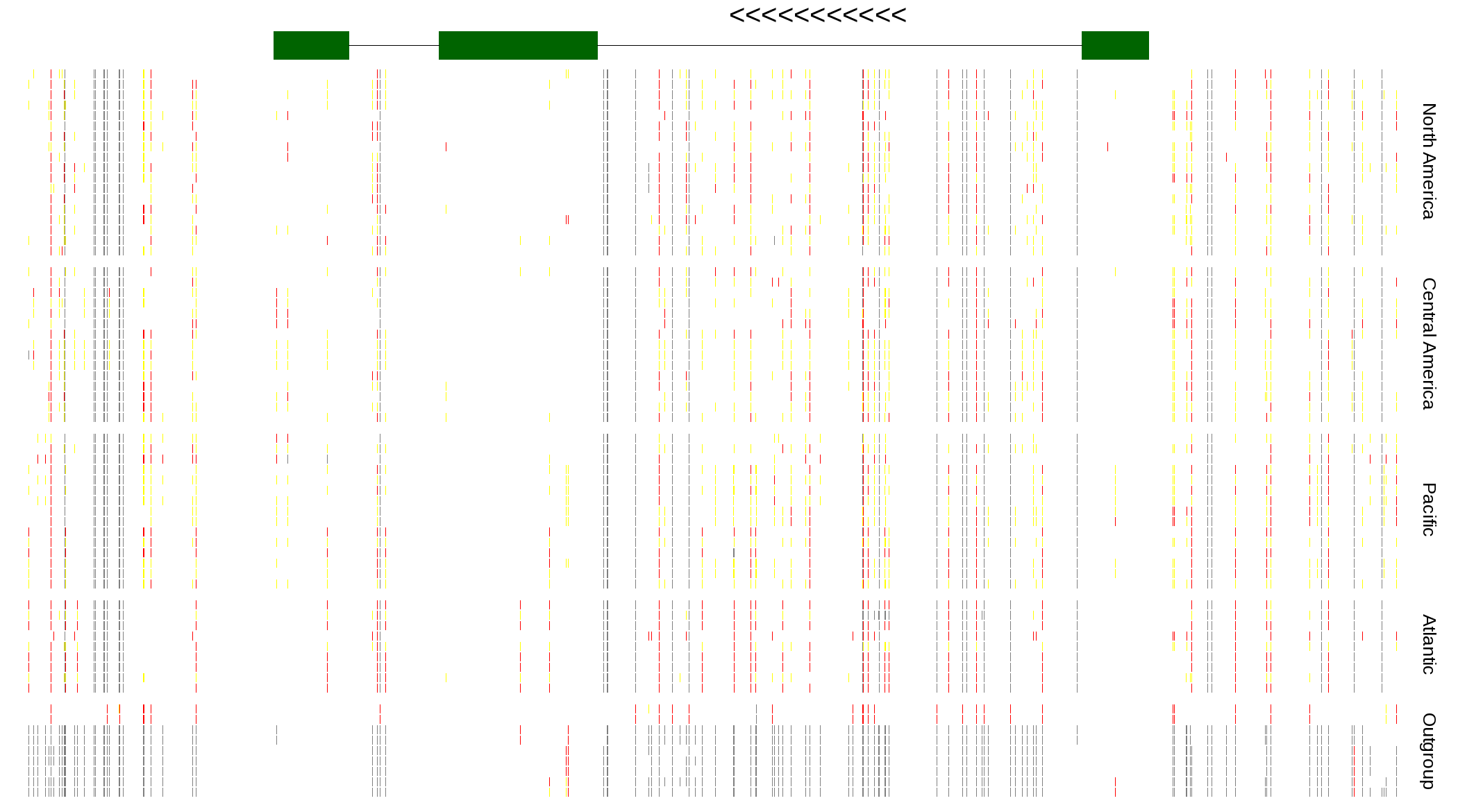
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS211437-TA
ATGCCTCTCCAGCTGACCAAACAAGCTCCTCAGTGGAAGACCACGGCCGTCGTCAACGGCGAGTTCAAGGACATCGCTCTGTCCGACTACAAGGGGAAATACGTTGTGCTGTTCTTCTACCCCCTGGACTTCACGTTCGTGTGCCCTACCGAGATCATCGCTTTCTCGGAGCGCGCGGACGAGTTCCGCAAGTTGGGATGTGAGGTGATCGCCGCCTCCACGGACTCGCACTTCACTCACCTGGCGTGGATCAACACGCCCAGGAAGCAGGGCGGTCTGGGACCCATGAACATCCCCATCATCAGCGACAAGTCGCACCGCATCGCCCGGGACTACGGCGTGCTCAACGAGGAGTCCGGGATACCCTTCAGGGGACTCTTCATCATCGACGACAAACAGAACCTCAGACAGGTCACCGTCAACGACCTGCCCGTGGGGAGGTCAGTAGACGAAACCCTTCGCCTGGTGCAGGCGTTCCAGTTCACTGACAAGCACGGCGAAGTGTGTCCGGCCAACTGGAGGCCGGGTGCCAAGACCATCAAACCTGACAGCAAGGCTGCCCAGGAGTACTTCGGAGACGTCAACTAA
>DPOGS211437-PA
MPLQLTKQAPQWKTTAVVNGEFKDIALSDYKGKYVVLFFYPLDFTFVCPTEIIAFSERADEFRKLGCEVIAASTDSHFTHLAWINTPRKQGGLGPMNIPIISDKSHRIARDYGVLNEESGIPFRGLFIIDDKQNLRQVTVNDLPVGRSVDETLRLVQAFQFTDKHGEVCPANWRPGAKTIKPDSKAAQEYFGDVN-