| DPOGS211614 | ||
|---|---|---|
| Transcript | DPOGS211614-TA | 573 bp |
| Protein | DPOGS211614-PA | 190 aa |
| Genomic position | DPSCF300232 + 208687-212761 | |
| RNAseq coverage | 100x (Rank: top 61%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL014013 | 1e-103 | 89.47% | |
| Bombyx | BGIBMGA008235-TA | 3e-55 | 75.76% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_E3WKY1 | 1e-57 | 55.44% | Putative uncharacterized protein n=13 Tax=Pancrustacea RepID=E3WKY1_ANODA |
| NCBI RefSeq | XP_001946608.1 | 6e-55 | 56.19% | PREDICTED: similar to AGAP012257-PA [Acyrthosiphon pisum] |
| NCBI nr blastp | gi|312385086 | 3e-57 | 55.44% | hypothetical protein AND_01190 [Anopheles darlingi] |
| NCBI nr blastx | gi|312385086 | 3e-56 | 55.44% | hypothetical protein AND_01190 [Anopheles darlingi] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0015671 | 1.2e-25 | oxygen transport | |
| GO:0020037 | 1.2e-25 | heme binding | ||
| GO:0005506 | 1.2e-25 | iron ion binding | ||
| GO:0019825 | 1.2e-25 | oxygen binding | ||
| KEGG pathway | ||||
| InterPro domain | [30-184] IPR009050 | 2.9e-27 | Globin-like | |
| [26-178] IPR012292 | 1.2e-25 | Globin | ||
| [41-143] IPR000971 | 1.8e-14 | Globin, subset | ||
| Orthology group | MCL15566 | Insect specific | ||
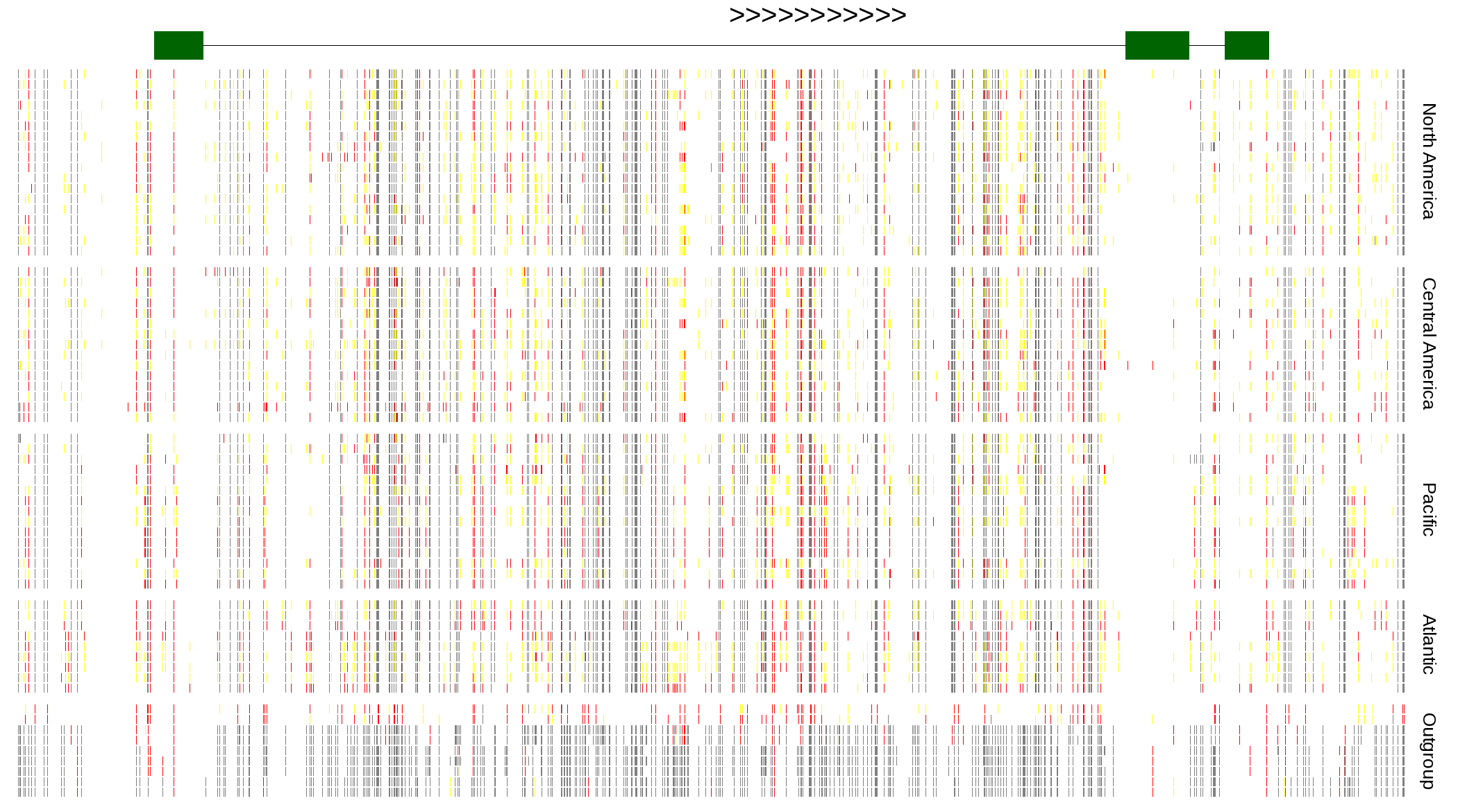
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS211614-TA
ATGGGTTGTAAATTGTCTCAGCTAGCATCTTCCGAATTTTCACACGATCCCTTCGAGAAACCTCCTCCTCCGTCCGATCCACGGTCACCTCTCACTGCCAAGCAGCAGTACTGCATGCTGGCCTCTTGGAAGGGAGTGTTCCGACAGGTCGAGAAAACTGGCATTCTCTTGTTTGTTAAATTGTTCGAAGAGAATGAAGAGCTTCTACATCTATTTGAAAAGTTCCGAGAGCTGCGCACCAAAGAAGCTATAGTCAGTTCTGCGGAACTGGCCGAGCACGCTACACAGGTGATGCACACACTAGACGAGGGCATCAAGGGGCTGGCGGATATGGACTCATTCTTCACTTACGTCAGACACGTGGGTGGCACGCATCGCCAGGTGCCCGGGTTTAAAGCAGAAAACTTTATGAAAATCGAGCAACCCTTCCTTGAAGCAGCCAAGACAACCTTGGGAGAGAGATACACCCCGAATATTGAAAATATATACAAGTTAACAATACGGTTCATTTTAGAGAATCTGGTCAAAGGTTACGAGGAGGCAGGGGAGGAAAACGGAACGACACAAACCTGA
>DPOGS211614-PA
MGCKLSQLASSEFSHDPFEKPPPPSDPRSPLTAKQQYCMLASWKGVFRQVEKTGILLFVKLFEENEELLHLFEKFRELRTKEAIVSSAELAEHATQVMHTLDEGIKGLADMDSFFTYVRHVGGTHRQVPGFKAENFMKIEQPFLEAAKTTLGERYTPNIENIYKLTIRFILENLVKGYEEAGEENGTTQT-