| DPOGS212194 | ||
|---|---|---|
| Transcript | DPOGS212194-TA | 357 bp |
| Protein | DPOGS212194-PA | 118 aa |
| Genomic position | DPSCF300459 + 123964-127724 | |
| RNAseq coverage | 494x (Rank: top 25%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL005525 | 7e-67 | 99.15% | |
| Bombyx | BGIBMGA001881-TA | 1e-48 | 100.00% | |
| Drosophila | Ih-PE | 4e-28 | 67.82% | |
| EBI UniRef50 | UniRef50_F4W9U8 | 5e-34 | 75.76% | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 n=1 Tax=Acromyrmex echinatior RepID=F4W9U8_ACREC |
| NCBI RefSeq | XP_001604549.1 | 2e-35 | 81.18% | PREDICTED: similar to hyperpolarization-activated ion channel isoform 1 [Nasonia vitripennis] |
| NCBI nr blastp | gi|3970750 | 3e-48 | 98.91% | cyclic nucleotide and voltage-activated ion channel [Heliothis virescens] |
| NCBI nr blastx | gi|3970750 | 2e-46 | 98.91% | cyclic nucleotide and voltage-activated ion channel [Heliothis virescens] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| InterPro domain | [73-118] IPR013621 | 3.1e-19 | Ion transport N-terminal | |
| Orthology group | MCL19720 | Insect specific | ||
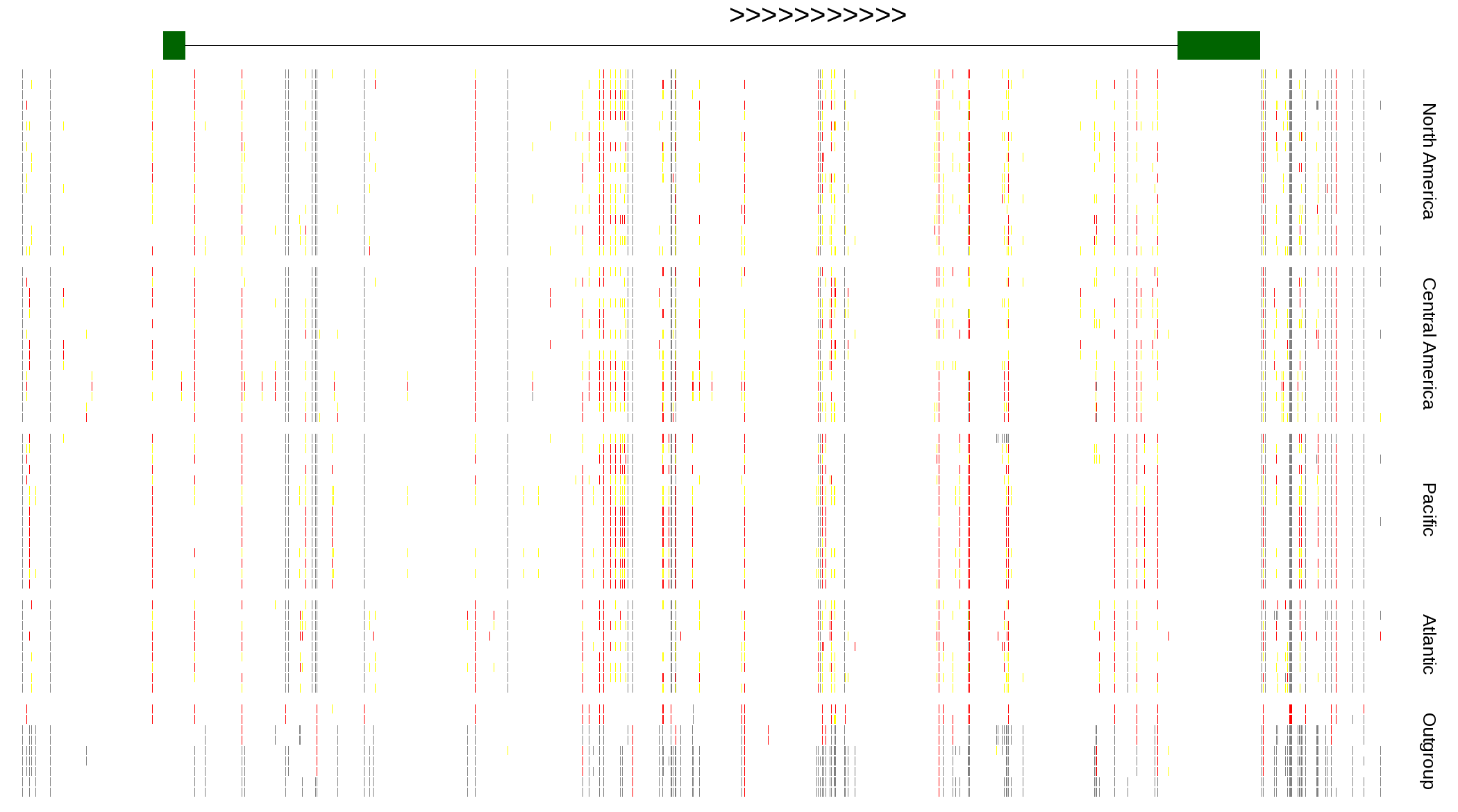
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS212194-TA
ATGTGCTGTTCAGGATTCACTAATTACTTGTTCCATAAACTAAGGAAATACACATGTGGGCTGGCGAGCCGCGAAAGCGACGCCGATATGTCGCTGAAGCAAGGCAGCGGCACCGGCAAAGTCCACTTCGGTGGTTTGGACGATGTGTCGTTGTACGGCACGCCAGTGGAGCCCGCGCCACCCGCACCCGACGCCAAGCAAGGATTCCTGCGAAACCAACTCCAAGCCCTCTTCCAGCCCACGGACAACAAGCTAGCCATGAAACTCTTCGGATCCAAGAAGGCACTCATGAAGGAGCGCATCAGACAGAAAGCGGCCGGTCACTGGGTTATACATCCGTGTAGTAGTTTTAGGTAA
>DPOGS212194-PA
MCCSGFTNYLFHKLRKYTCGLASRESDADMSLKQGSGTGKVHFGGLDDVSLYGTPVEPAPPAPDAKQGFLRNQLQALFQPTDNKLAMKLFGSKKALMKERIRQKAAGHWVIHPCSSFR-