| DPOGS213030 | ||
|---|---|---|
| Transcript | DPOGS213030-TA | 666 bp |
| Protein | DPOGS213030-PA | 221 aa |
| Genomic position | DPSCF300024 + 401422-402633 | |
| RNAseq coverage | 202x (Rank: top 47%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL020888 | 1e-87 | 69.59% | |
| Bombyx | BGIBMGA002881-TA | 8e-25 | 31.58% | |
| Drosophila | slv-PA | 4e-42 | 37.96% | |
| EBI UniRef50 | UniRef50_B4NMK1 | 4e-41 | 40.28% | GK23163 n=2 Tax=Sophophora RepID=B4NMK1_DROWI |
| NCBI RefSeq | NP_001161261.1 | 2e-45 | 41.83% | recombination activating gene 1 activating protein 1 [Nasonia vitripennis] |
| NCBI nr blastp | gi|268370163 | 4e-44 | 41.83% | recombination activating gene 1 activating protein 1 [Nasonia vitripennis] |
| NCBI nr blastx | gi|268370163 | 3e-46 | 41.83% | recombination activating gene 1 activating protein 1 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016021 | 4.4e-51 | integral to membrane | |
| GO:0006310 | 4.4e-51 | DNA recombination | ||
| KEGG pathway | ||||
| InterPro domain | [13-211] IPR018180 | 4.4e-51 | RAG1-activating protein 1 homologue, Rag1ap1 | |
| [13-211] IPR004316 | 4.4e-51 | RAG1-activating protein-1-related | ||
| Orthology group | MCL14924 | Single-copy universal gene | ||
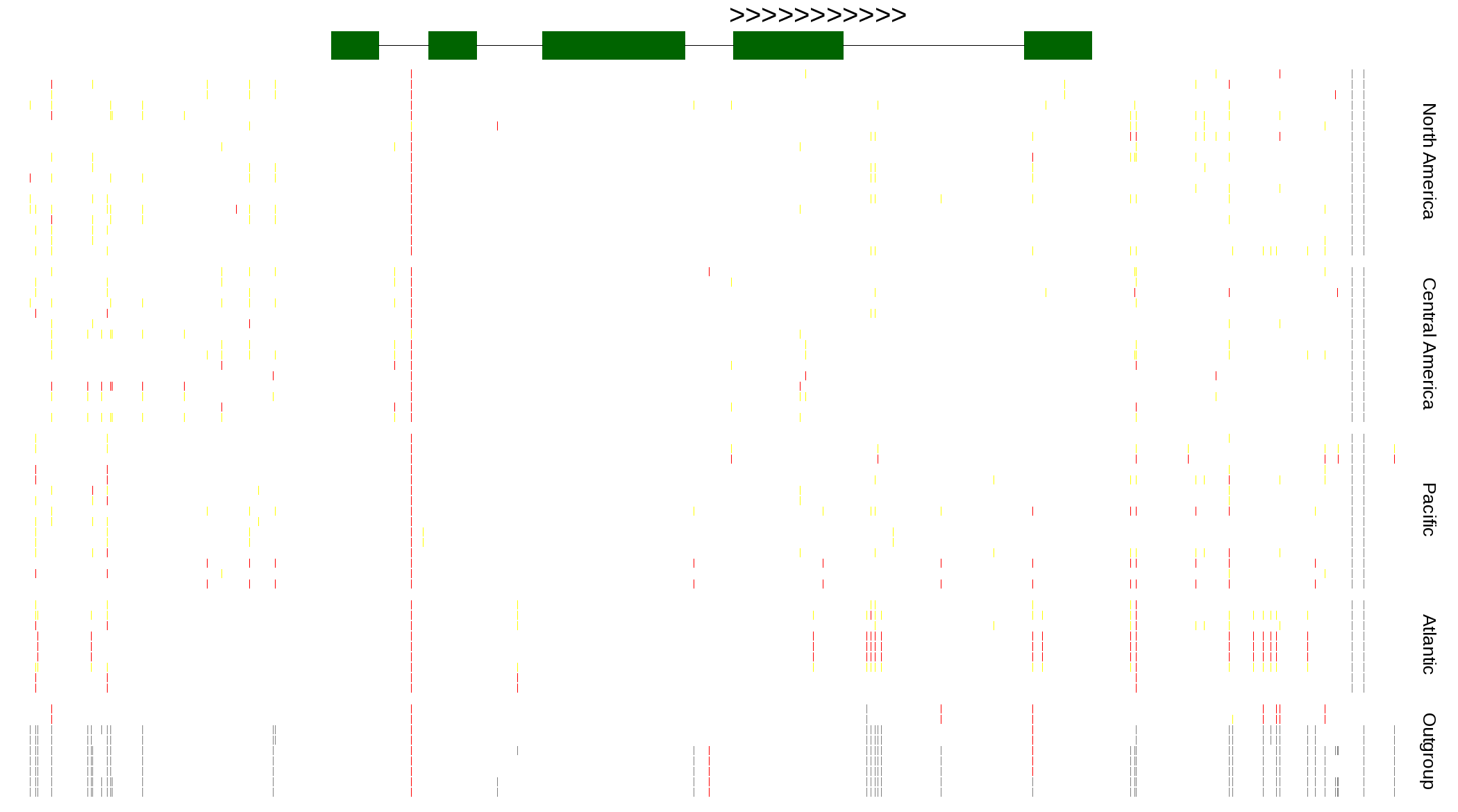
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS213030-TA
ATGTTTTTAACAAATTTTAATGAATTCGTTGCTAATTTGGCTGTTATTACAACAATAATACAATTTTTGTCGGGGATATTGGTTTGTCGACAGTATGTAGTTAATCGCACAACAGCAGAAGCTTCACCACTACCATTTATTTGCGGTTTTCTATCTTCTGGTTTATGGCTTTTATATGGGATATGTAAGCCTGATAGTAAAATAATAATAGTTAATGTTGTTGGTGTTTTGCTGATGCTATCATACAGTATAGTATTCTATGTTTACACTTTTAAAAAGTCATCAGTATTAAAACAGTCTCTAGTCGCAATAATATTATATCTAGTAATGGTTGTTTATATGTCTACTGAAATTGACAATGAAATTCTACTGGTTAGATTAGGTTATTCTGCTTGCCTCTTAACATTGTTAACTATAAGTGCACCAATGAGCAAGTTATTCTATGTCATTAGAACAAAATGCACTGATTGCCTTCCATTCCCAATGATTTTTATGTCATTTATTGTGTCCAGTCTATGGTTTATCTATGGTTGTATAGTACAAGATGTATTTTTATCGATTCCCAACTTCATAGGTGCTTCTTTGGCTGTGGCACAGTTGTCACTATTTGTCGTCTACCCGAGTGTGCCACAAACACCTCTGTTATTAAAGATGACAGAGGCATAA
>DPOGS213030-PA
MFLTNFNEFVANLAVITTIIQFLSGILVCRQYVVNRTTAEASPLPFICGFLSSGLWLLYGICKPDSKIIIVNVVGVLLMLSYSIVFYVYTFKKSSVLKQSLVAIILYLVMVVYMSTEIDNEILLVRLGYSACLLTLLTISAPMSKLFYVIRTKCTDCLPFPMIFMSFIVSSLWFIYGCIVQDVFLSIPNFIGASLAVAQLSLFVVYPSVPQTPLLLKMTEA-