| DPOGS213073 | ||
|---|---|---|
| Transcript | DPOGS213073-TA | 615 bp |
| Protein | DPOGS213073-PA | 204 aa |
| Genomic position | DPSCF300016 - 555772-557091 | |
| RNAseq coverage | 445x (Rank: top 28%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004014 | 2e-71 | 60.91% | |
| Bombyx | BGIBMGA007690-TA | 3e-114 | 94.53% | |
| Drosophila | PpV-PA | 8e-93 | 75.25% | |
| EBI UniRef50 | UniRef50_Q9CQR6 | 8e-92 | 78.61% | Serine/threonine-protein phosphatase 6 catalytic subunit n=23 Tax=Euteleostomi RepID=PPP6_MOUSE |
| NCBI RefSeq | NP_001040439.1 | 8e-113 | 94.53% | serine/threonine protein phosphatase 6 [Bombyx mori] |
| NCBI nr blastp | gi|114051936 | 1e-111 | 94.53% | serine/threonine protein phosphatase 6 [Bombyx mori] |
| NCBI nr blastx | gi|114051936 | 8e-115 | 94.53% | serine/threonine protein phosphatase 6 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016787 | 2.3e-67 | hydrolase activity | |
| KEGG pathway | ||||
| InterPro domain | [46-73] IPR006186 | 2.3e-67 | Serine/threonine-specific protein phosphatase/bis(5-nucleosyl)-tetraphosphatase | |
| [46-175] IPR004843 | 6.6e-38 | Metallophosphoesterase domain | ||
| Orthology group | MCL14801 | Single-copy universal gene | ||
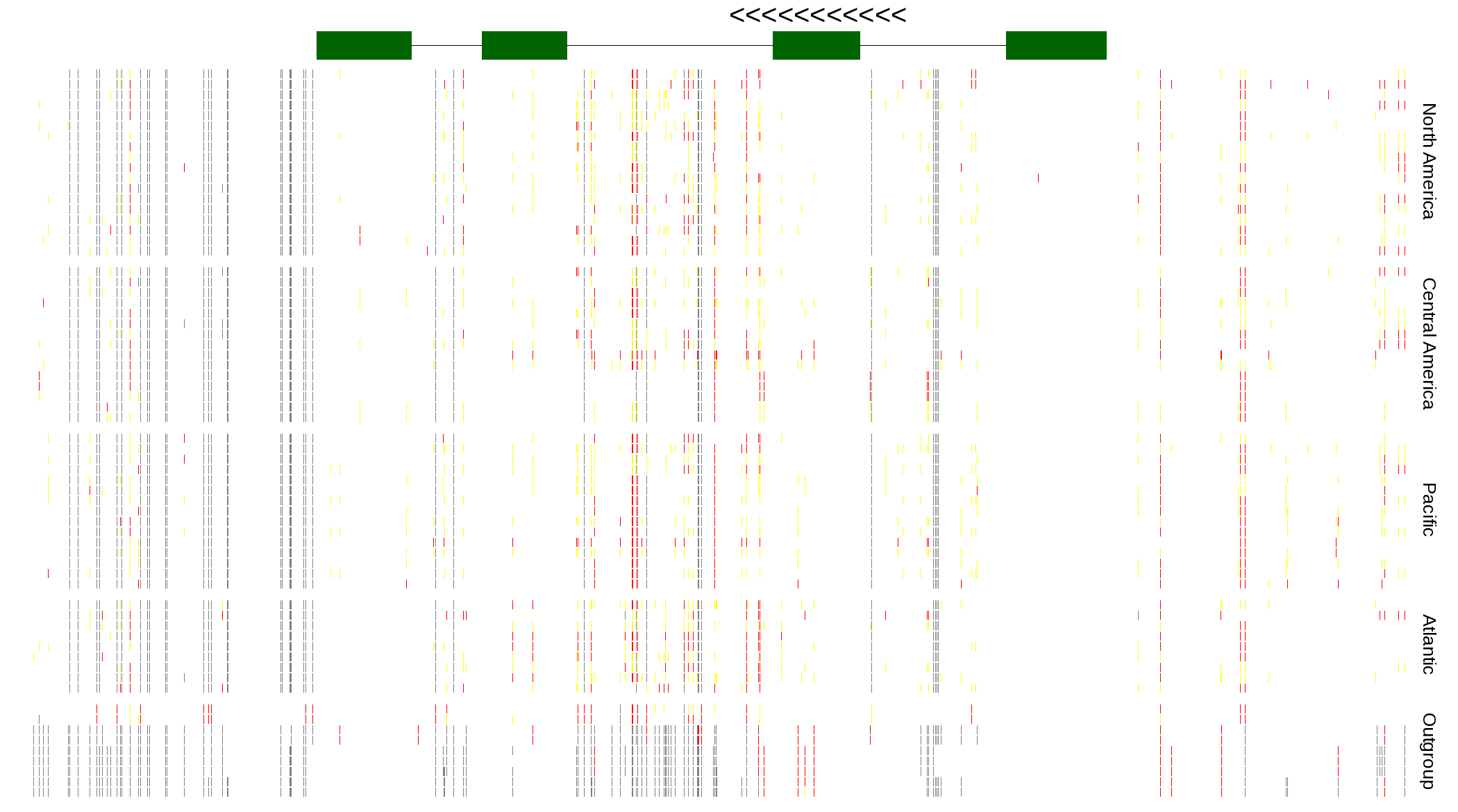
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS213073-TA
ATGACGGGAGATGTTGATAAGTGGATAGAGATAGCAAAGAGGTGTAAATACTTGCCGGAAGATGATCTGAGAGAACTCTGTAATATAGTATGTGATTTATTACTAGAAGAACCGAATGTTCAACCAGTCCAAACACCAGTTACTGTTTGTGGTGATATTCATGGTCAGTTCTATGACTTAGAAGAGCTATTCCATATAGGAGGACAAGTTCCTAGCACTCAGTATATATTTATGGGAGATTATGTCGATCGAGGGTACTACAGTCTTGAAACGTTGACACTCTTAATGGCTCTTAAAGCCAGGTACCCTGACAGGATAGTATTACTAAGAGGGAACCATGAGACCTGTCAAATAACCAAAGTTTATGGTTTCTATGATGAGTGTCTGAATAAATATGGAAATGCAAATGCATGGAAGGATTGTTGCAGGGTGTTTGACCTACTTACTGTAGCCGCGTTAATAGATGAGACAATTTTGTGTGTCCATGGAGGTCTGTCACCTGATATAGCAATGCTGGATCAAATCCGTTGTATTAATAGAAACCAACAGATTCCTCACAAAGGTGCCTTTTGTGACTTACTCTGGTCCGACCCAGCGGATGTCAAAACATGGTAA
>DPOGS213073-PA
MTGDVDKWIEIAKRCKYLPEDDLRELCNIVCDLLLEEPNVQPVQTPVTVCGDIHGQFYDLEELFHIGGQVPSTQYIFMGDYVDRGYYSLETLTLLMALKARYPDRIVLLRGNHETCQITKVYGFYDECLNKYGNANAWKDCCRVFDLLTVAALIDETILCVHGGLSPDIAMLDQIRCINRNQQIPHKGAFCDLLWSDPADVKTW-