| DPOGS213095 | ||
|---|---|---|
| Transcript | DPOGS213095-TA | 462 bp |
| Protein | DPOGS213095-PA | 153 aa |
| Genomic position | DPSCF300016 - 129637-130098 | |
| RNAseq coverage | 13564x (Rank: top 1%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006549 | 2e-78 | 88.24% | |
| Bombyx | BGIBMGA007701-TA | 7e-80 | 91.50% | |
| Drosophila | awd-PA | 1e-74 | 82.35% | |
| EBI UniRef50 | UniRef50_B0WCA1 | 5e-68 | 78.95% | Nucleoside diphosphate kinase n=4 Tax=Coelomata RepID=B0WCA1_CULQU |
| NCBI RefSeq | NP_001093284.1 | 2e-78 | 91.50% | abnormal wing disc-like protein [Bombyx mori] |
| NCBI nr blastp | gi|21435082 | 2e-79 | 92.16% | abnormal wing disc-like protein [Choristoneura parallela] |
| NCBI nr blastx | gi|21435082 | 6e-80 | 92.16% | abnormal wing disc-like protein [Choristoneura parallela] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006228 | 4.6e-107 | UTP biosynthetic process | |
| GO:0005524 | 4.6e-107 | ATP binding | ||
| GO:0006241 | 4.6e-107 | CTP biosynthetic process | ||
| GO:0006165 | 4.6e-107 | nucleoside diphosphate phosphorylation | ||
| GO:0004550 | 4.6e-107 | nucleoside diphosphate kinase activity | ||
| GO:0006183 | 4.6e-107 | GTP biosynthetic process | ||
| KEGG pathway | tca:655847 | 1e-73 | ||
| K00940 (E2.7.4.6, ndk) | maps-> | Purine metabolism | ||
| Pyrimidine metabolism | ||||
| InterPro domain | [4-153] IPR001564 | 4.6e-107 | Nucleoside diphosphate kinase | |
| Orthology group | MCL11416 | Multiple-copy universal gene | ||
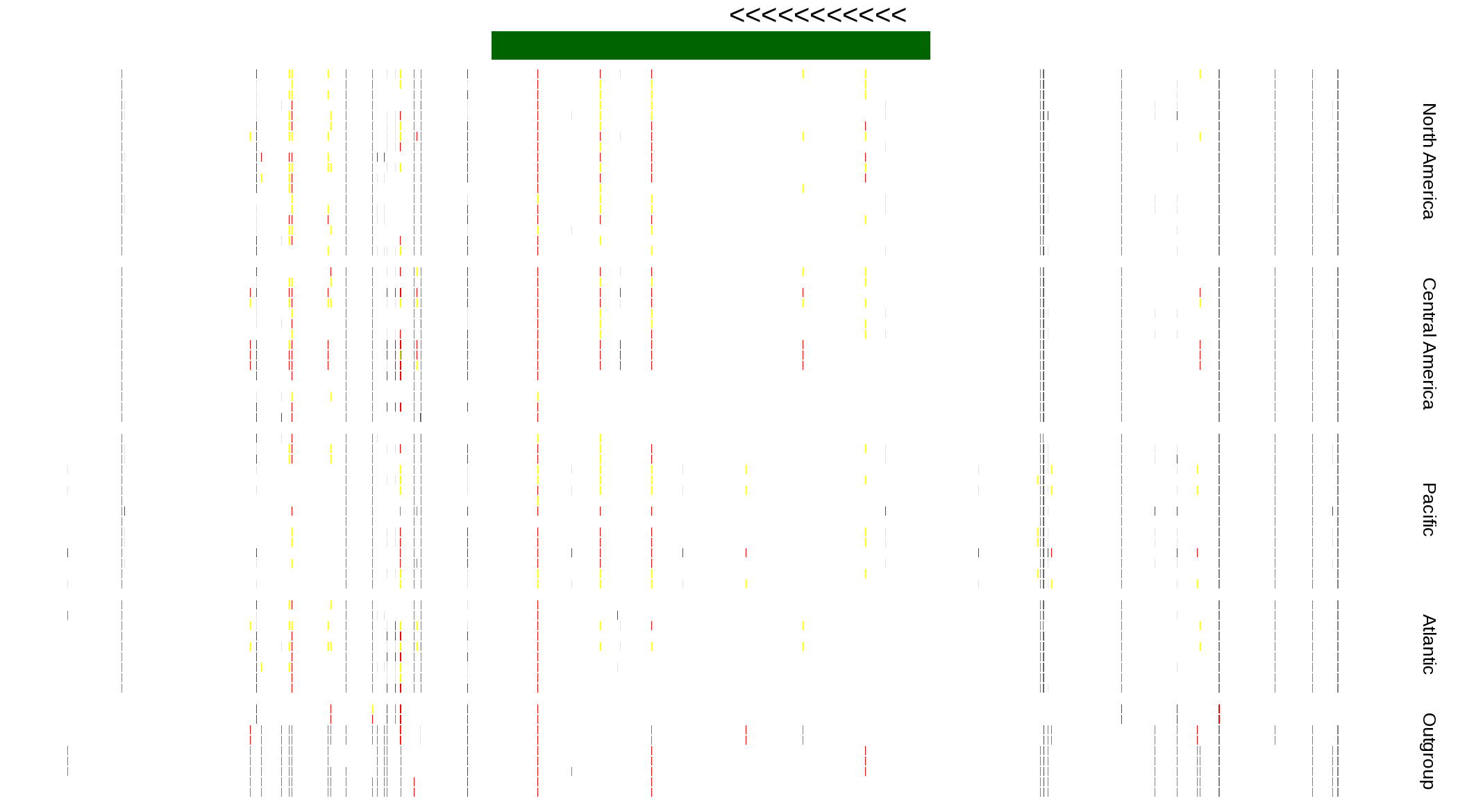
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS213095-TA
ATGGCTGAACCACGCGAAAGAACTTTCATCATGGTTAAACCAGACGGCGTACAAAGAGGACTCGTCGGCCAAATTATGGAAAGGTTCGAAAAAAAAGGTTTTAAGCTTGTAGGTCTTAAGTTTGTGTGGCCATCTGAAGAGCTTTTACAAAAGCATTACAGCGACCTGGCATCGCGACCATTCTTCCCTGGTCTTGTGAAATACATGAGTTCCGGGCCTGTTGTACCCATGGTTTGGGAAGGACTTAACGTAGTGAAGACCGGTCGTCAAATGCTCGGCGCAACAAACCCAGCTGACTCTTTACCTGGTACCATTCGTGGAGATCTGTGCATTGAAGTTGGCCGCAACATTATTCACGGCTCTGACAGTGTTGACTCAGCTAACAAGGAAATCAATCTCTGGTTCACTGAAAAAGAACTCGTCGGATGGACCCCTGCTGCCGAGAAATGGGTCTATGAATAA
>DPOGS213095-PA
MAEPRERTFIMVKPDGVQRGLVGQIMERFEKKGFKLVGLKFVWPSEELLQKHYSDLASRPFFPGLVKYMSSGPVVPMVWEGLNVVKTGRQMLGATNPADSLPGTIRGDLCIEVGRNIIHGSDSVDSANKEINLWFTEKELVGWTPAAEKWVYE-