| DPOGS214007 | ||
|---|---|---|
| Transcript | DPOGS214007-TA | 618 bp |
| Protein | DPOGS214007-PA | 205 aa |
| Genomic position | DPSCF300313 - 32661-34395 | |
| RNAseq coverage | 2362x (Rank: top 5%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL013815 | 2e-103 | 84.88% | |
| Bombyx | BGIBMGA004059-TA | 1e-95 | 83.33% | |
| Drosophila | CG2852-PA | 7e-81 | 77.72% | |
| EBI UniRef50 | UniRef50_E4XG60 | 9e-79 | 78.69% | Peptidyl-prolyl cis-trans isomerase n=5 Tax=Bilateria RepID=E4XG60_OIKDI |
| NCBI RefSeq | NP_001040479.1 | 3e-96 | 82.44% | peptidylprolyl isomerase B [Bombyx mori] |
| NCBI nr blastp | gi|114052472 | 4e-95 | 82.44% | peptidylprolyl isomerase B precursor [Bombyx mori] |
| NCBI nr blastx | gi|114052472 | 3e-95 | 82.84% | peptidylprolyl isomerase B precursor [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006457 | 4e-74 | protein folding | |
| GO:0003755 | 4e-74 | peptidyl-prolyl cis-trans isomerase activity | ||
| KEGG pathway | ||||
| InterPro domain | [27-188] IPR002130 | 4e-74 | Peptidyl-prolyl cis-trans isomerase, cyclophilin-type | |
| [14-190] IPR015891 | 3.2e-70 | Cyclophilin-like | ||
| Orthology group | MCL13452 | Single-copy universal gene | ||
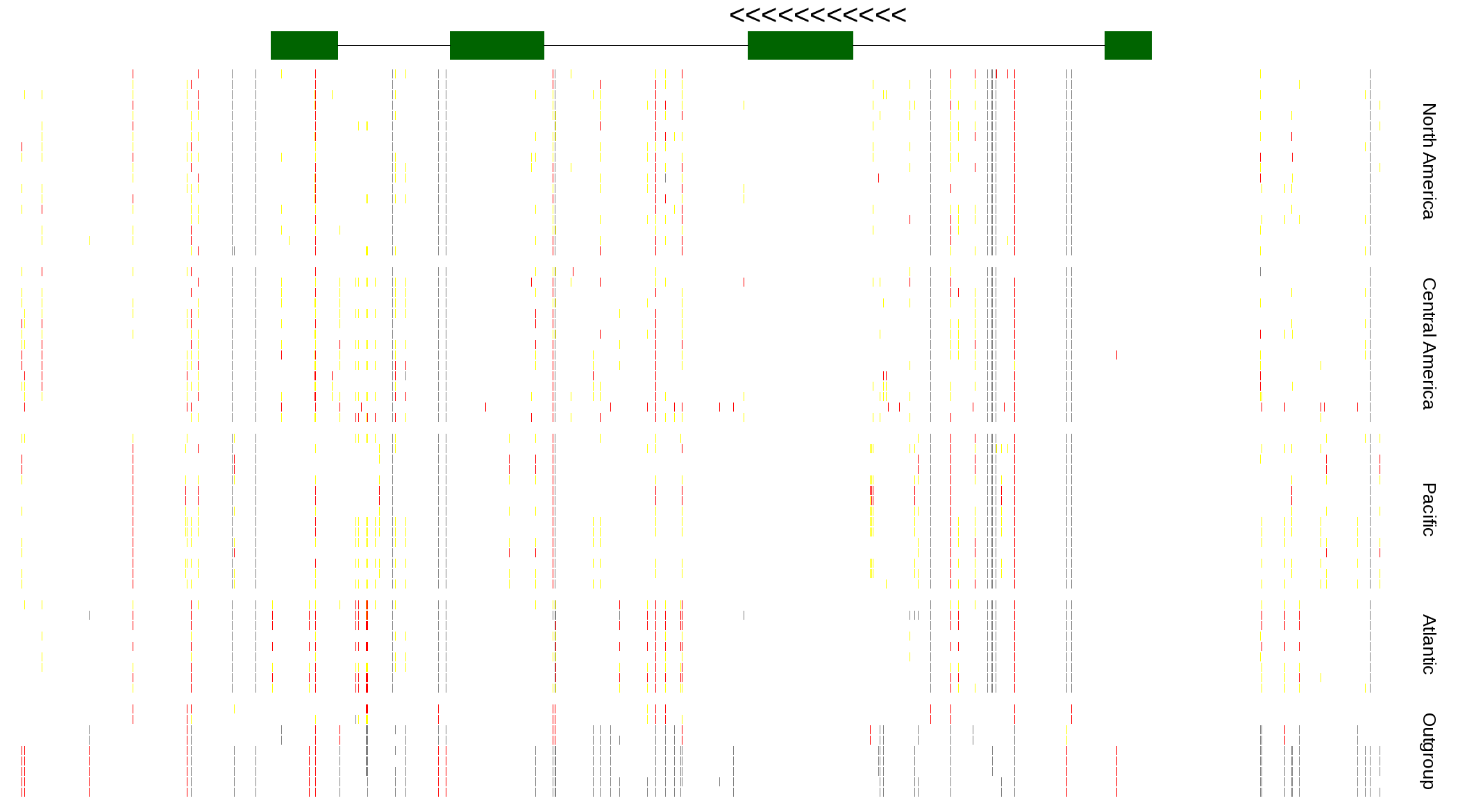
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214007-TA
ATGGGTCTCTTAACGGCGACTCTTGGAATTCTATTCCTGGTGTCAACTGTCTGGTCAGCAGATGAACCCAAAGGACCCAAAGTTACCCACAAGGTAAAATTTGACATGAGCATTGGTGGTGACCCGATAGGAACTATTGTAATTGGTTTGTTTGGAAAAACAGTTCCTAAGACTGTAGAGAATTTCTACCAACTTGCCCAAAAGCCTGAGGGTCAAGGTTACAAAGGTAGCAAATTCCACAGGGTCATTGATAATTTCATGATTCAAGGTGGTGACTTCACCAAGGGAGATGGAACTGGAGGCATGAGCATTTATGGAGAGAGGTTTGAGGATGAAAACTTCAAGCTCCGTCACTATGGTGCTGGATGGTTGTCAATGGCCAATGCTGGTAAGGACACCAATGGCTCTCAGTTCTTCATCACCACCACCAAAACTCCCTGGTTAGATGGCAGACATGTTGTCTTTGGAAAAATTCTTGAAGGAATGGATGTGGTGCGTAAGATTGAAAAGACGGTAACTGGAGCAAATGACCGGCCAGTTAAGGATGTCGTAATAACAAACTCATCAGCGGAAGTTGTATCAGAACCTTTTGCTGTATCAAAAGAAAGTGCTCAATGA
>DPOGS214007-PA
MGLLTATLGILFLVSTVWSADEPKGPKVTHKVKFDMSIGGDPIGTIVIGLFGKTVPKTVENFYQLAQKPEGQGYKGSKFHRVIDNFMIQGGDFTKGDGTGGMSIYGERFEDENFKLRHYGAGWLSMANAGKDTNGSQFFITTTKTPWLDGRHVVFGKILEGMDVVRKIEKTVTGANDRPVKDVVITNSSAEVVSEPFAVSKESAQ-