| DPOGS214154 | ||
|---|---|---|
| Transcript | DPOGS214154-TA | 552 bp |
| Protein | DPOGS214154-PA | 183 aa |
| Genomic position | DPSCF300014 - 754226-755158 | |
| RNAseq coverage | 852x (Rank: top 15%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL002248 | 1e-99 | 93.44% | |
| Bombyx | BGIBMGA006200-TA | 3e-100 | 92.90% | |
| Drosophila | CG10470-PA | 1e-91 | 85.25% | |
| EBI UniRef50 | UniRef50_Q9UM00 | 6e-71 | 74.32% | Transmembrane and coiled-coil domain-containing protein 1 n=65 Tax=Bilateria RepID=TMCO1_HUMAN |
| NCBI RefSeq | XP_970659.1 | 4e-94 | 87.98% | PREDICTED: similar to CG10470 CG10470-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|91082035 | 6e-93 | 87.98% | PREDICTED: similar to CG10470 CG10470-PA [Tribolium castaneum] |
| NCBI nr blastx | gi|307189349 | 1e-89 | 87.43% | Transmembrane and coiled-coil domains protein 1 [Camponotus floridanus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016020 | 5.2e-147 | membrane | |
| KEGG pathway | ||||
| InterPro domain | [1-183] IPR008559 | 5.2e-147 | Uncharacterised conserved protein UCP023322, transmembrane eukaryotic | |
| [7-163] IPR002809 | 1.8e-46 | Protein of unknown function DUF106, transmembrane | ||
| Orthology group | MCL12093 | Single-copy universal gene | ||
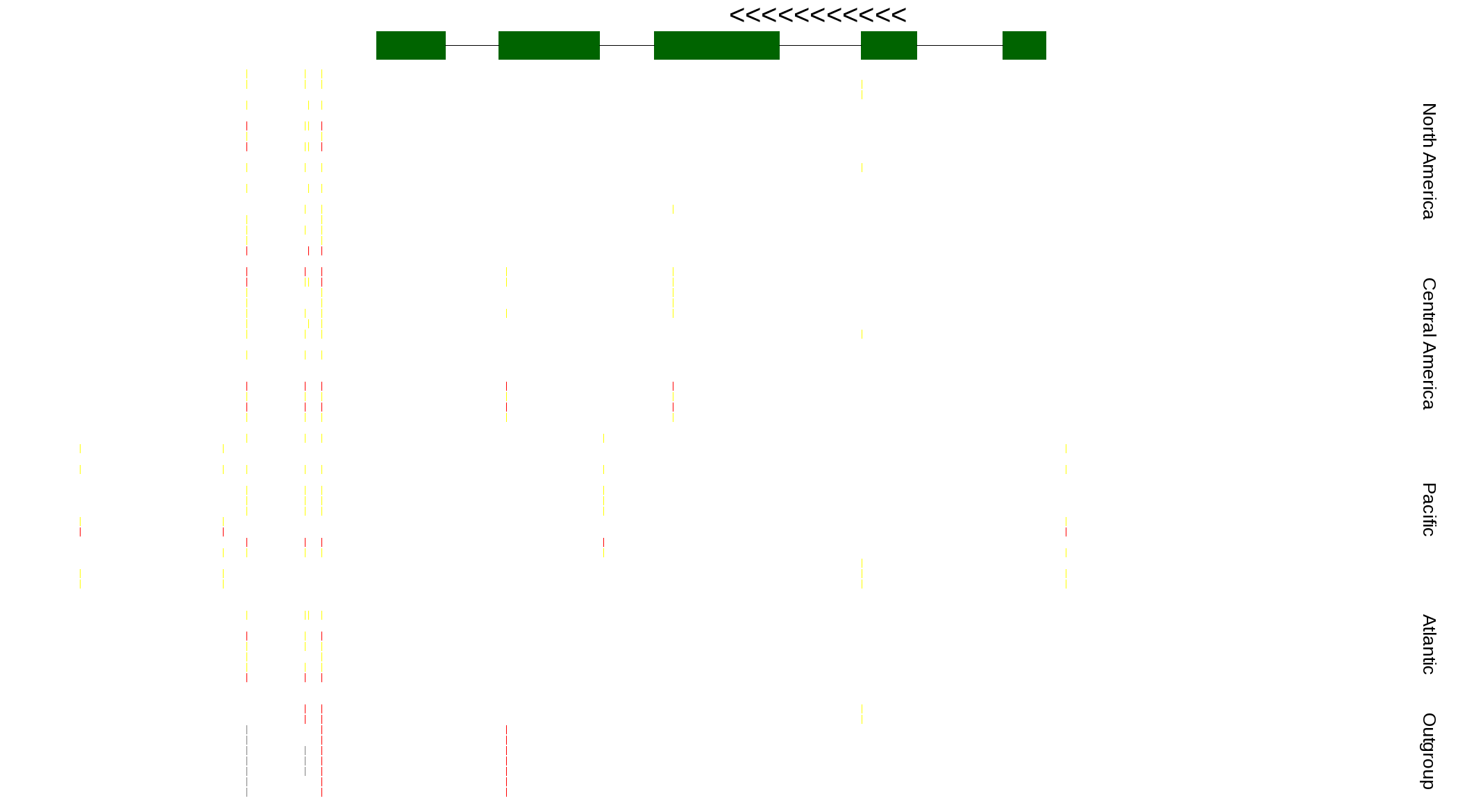
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214154-TA
ATGTGGGGTGATCCTTTATTGATAGTATTTATTTCAGTATGTACTGCTTTCTTGGGAGAAGGACTTACATGGATATTGGTATATAGAACTGAAAAATACCAAAAATTAAAGGTAGAAGTTGAGCGCCAAAGTAAGAAGTTGGAGAAAAGAAAAGAAGCACATGGTGATTCTTTGGATAAGCAACATAAGAAAAAAATTGAACGTGAAGAGGAGAGGCTGAAAAATAATAATAGGGACCTGTCCCTTGTTAAAATGAAATCCATGTTTGCCATTGGTTTTGCATTTACAGCTTTACTGAGCATGTTTAATAACATATTTGATGGTAGAGTGGTAGCAAAGCTTCCTTTTCACCCAATATCCTGGATCCAAGGTCTCAGTCACCGTAATTTACCTGGAGATGACTTCACAGATTGCTCATTCATTTTTCTGTACATTTTATGCACTATGAGCATAAGGCAAAATATTCAGAAGCTACTTGGATTTGCACCTTCTCGTGCAGCATCTCAACAAGGCGGAGCCCTTTTTGCAACTCCTCCAGCTCAGTTTAAATAA
>DPOGS214154-PA
MWGDPLLIVFISVCTAFLGEGLTWILVYRTEKYQKLKVEVERQSKKLEKRKEAHGDSLDKQHKKKIEREEERLKNNNRDLSLVKMKSMFAIGFAFTALLSMFNNIFDGRVVAKLPFHPISWIQGLSHRNLPGDDFTDCSFIFLYILCTMSIRQNIQKLLGFAPSRAASQQGGALFATPPAQFK-