| DPOGS214447 | ||
|---|---|---|
| Transcript | DPOGS214447-TA | 627 bp |
| Protein | DPOGS214447-PA | 208 aa |
| Genomic position | DPSCF300441 - 67125-70447 | |
| RNAseq coverage | 5487x (Rank: top 2%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004430 | 2e-96 | 77.78% | |
| Bombyx | BGIBMGA009614-TA | 6e-87 | 70.67% | |
| Drosophila | CG9796-PA | 4e-32 | 36.55% | |
| EBI UniRef50 | UniRef50_Q2F5Z1 | 4e-81 | 64.76% | Lysosomal thiol reductase IP30 isoform 1 n=2 Tax=Bombyx mori RepID=Q2F5Z1_BOMMO |
| NCBI RefSeq | NP_001103767.1 | 2e-84 | 70.19% | lysosomal thiol reductase IP30 isoform 2 [Bombyx mori] |
| NCBI nr blastp | gi|160333474 | 4e-83 | 70.19% | lysosomal thiol reductase IP30 isoform 2 precursor [Bombyx mori] |
| NCBI nr blastx | gi|160333474 | 4e-82 | 70.19% | lysosomal thiol reductase IP30 isoform 2 precursor [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | dre:336503 | 2e-29 | ||
| K08059 (IFI30, GILT) | maps-> | Antigen processing and presentation | ||
| InterPro domain | [24-198] IPR004911 | 5.5e-60 | Gamma interferon inducible lysosomal thiol reductase GILT | |
| Orthology group | MCL10930 | Multiple-copy universal gene | ||
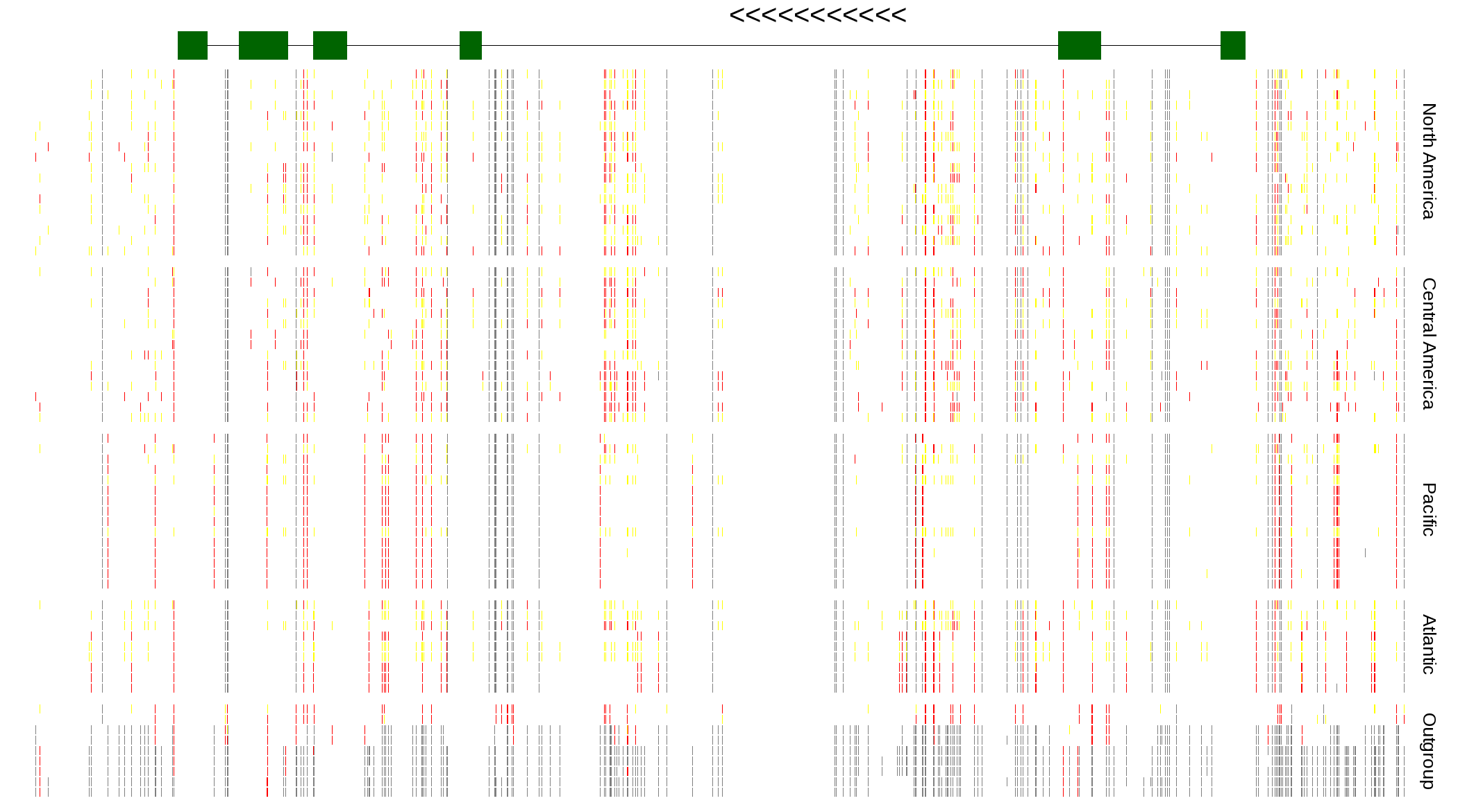
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214447-TA
ATGCGATCTCTGCTTGTTTTATTTGTGCTGACCGTCCTTTGTGGAGTTCACGCGAAGGGAAAACATGACGACAAGGTGAAGATAGCTGTGTACTACGAGTCACTATGCCCGGACAGTAAGAAGTTCATAACGAGCCAGCTGGCCCCGGTCTGGAGAGACTTCCGGGGACAGGTCAAGGTCAAGATGGTGCCCTACGGGAAGGCCACCCACGACAAAGTGAACGGTAAATGGCAGTTCATTTGTCACCACGGACCCGACGAGTGTTACGGTAACAAGCTCCAGTCATGTATCCTGAAAGACCGCACGCTCCAGGACACCGACAAGATGGAACTCATCATCTGTCTCATGGGACAGGGGCATCCCGACAAGGCCATCGACACCTGTCTTGGTCAAGTCAAGAGAGAAGCCGAGTCCGCCAAGTTGAAGAAATGTGCGTCAGGGGCCCAGGGTGACGCGTTGCTGGCGTCCTACGGTGACAAGACGGACGTCGTGCAGAGACCGCTGTCCTTCGTGCCGACTGTTGTTATCAACGAGAAATTCGACCAAGCCGTCCAGGACGAGGCCGTGAGCGACCTGAAGGCGGTGGTGTGTCGCGTGGCCGCCAACAAACCAGCTCTCTGTCTTTAA
>DPOGS214447-PA
MRSLLVLFVLTVLCGVHAKGKHDDKVKIAVYYESLCPDSKKFITSQLAPVWRDFRGQVKVKMVPYGKATHDKVNGKWQFICHHGPDECYGNKLQSCILKDRTLQDTDKMELIICLMGQGHPDKAIDTCLGQVKREAESAKLKKCASGAQGDALLASYGDKTDVVQRPLSFVPTVVINEKFDQAVQDEAVSDLKAVVCRVAANKPALCL-