| DPOGS214468 | ||
|---|---|---|
| Transcript | DPOGS214468-TA | 393 bp |
| Protein | DPOGS214468-PA | 130 aa |
| Genomic position | DPSCF300122 - 625551-627392 | |
| RNAseq coverage | 80x (Rank: top 64%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL003746 | 2e-60 | 96.46% | |
| Bombyx | BGIBMGA013354-TA | 3e-30 | 77.78% | |
| Drosophila | onecut-PA | 2e-38 | 85.54% | |
| EBI UniRef50 | UniRef50_Q9NJB5 | 3e-36 | 85.54% | Homeobox protein onecut n=7 Tax=melanogaster subgroup RepID=ONEC_DROME |
| NCBI RefSeq | NP_524842.2 | 4e-37 | 85.54% | onecut [Drosophila melanogaster] |
| NCBI nr blastp | gi|350426167 | 2e-37 | 80.00% | PREDICTED: hepatocyte nuclear factor 6-like [Bombus impatiens] |
| NCBI nr blastx | gi|350426167 | 3e-36 | 80.00% | PREDICTED: hepatocyte nuclear factor 6-like [Bombus impatiens] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003677 | 2.8e-17 | DNA binding | |
| GO:0006355 | 2.8e-17 | regulation of transcription, DNA-dependent | ||
| GO:0043565 | 1.5e-13 | sequence-specific DNA binding | ||
| GO:0003700 | 1.5e-13 | sequence-specific DNA binding transcription factor activity | ||
| GO:0005515 | 3.3e-13 | protein binding | ||
| KEGG pathway | xtr:100101763 | 6e-27 | ||
| K08026 (ONECUT1, HNF6) | maps-> | Maturity onset diabetes of the young | ||
| InterPro domain | [15-81] IPR012287 | 2.8e-17 | Homeodomain-related | |
| [29-91] IPR001356 | 1.5e-13 | Homeobox | ||
| [28-96] IPR009057 | 3.3e-13 | Homeodomain-like | ||
| Orthology group | MCL26801 | Insect specific | ||
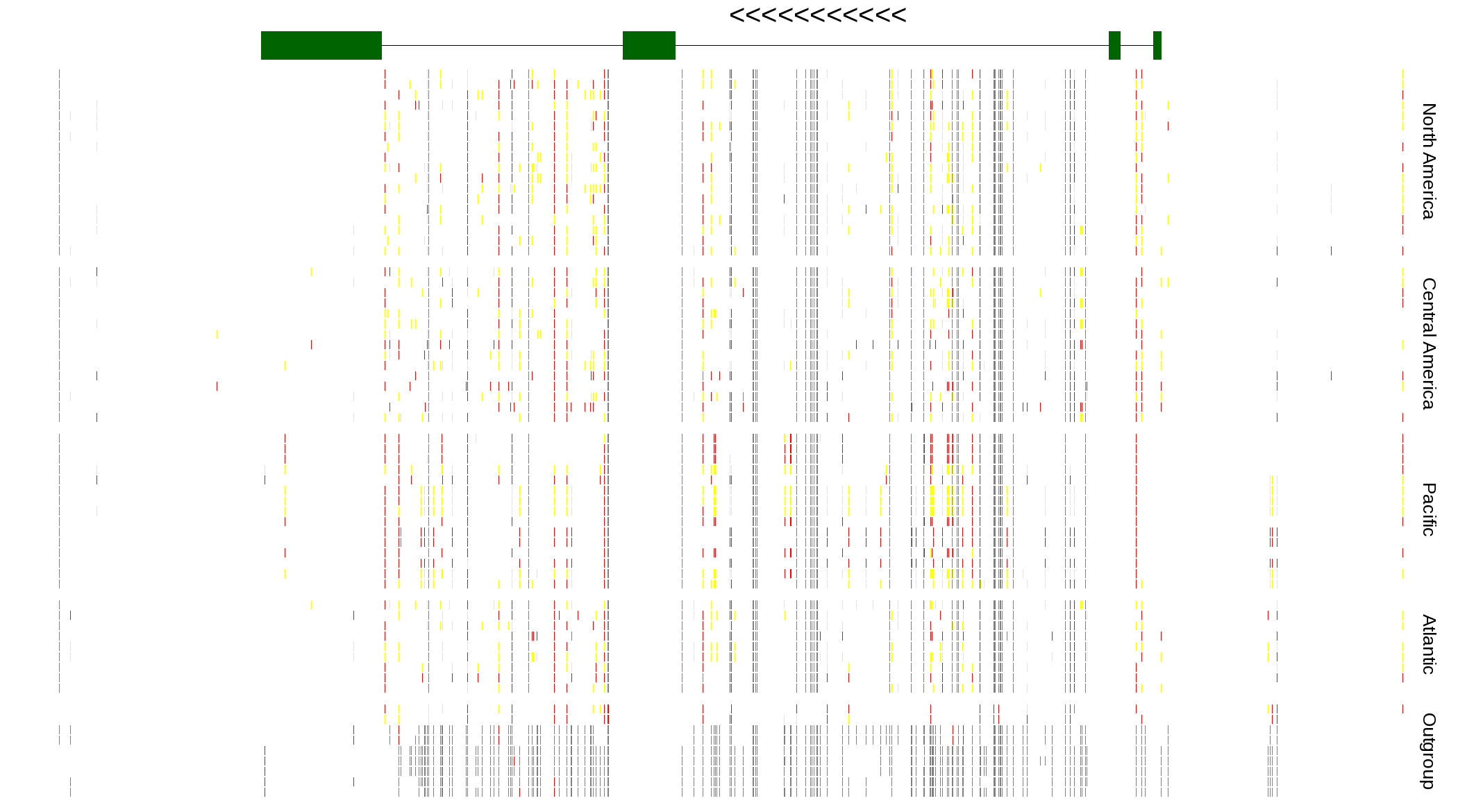
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214468-TA
ATGACTGATGCAAGGACGGCCCAGATACCACAGAGAGGCAGCTGCAAGCGCAAGGAGGATATGGCCTCGGATACCCTTCCTTCCCCGAAGAAGCCGCGTTTGGTTTTCACAGATCTGCAGCGGCGTACTCTGCAGGCTATTTTTAAGGAAACAAAGAGACCATCAAAAGAGATGCAAGTAACGATAGCGCGTCAGCTCGGTCTAGAGCCGACCACTGTTGGCAACTTCTTCATGAACGCTCGCAGACGATCCATGGACAAATGGAAGGATGACGACGCACCATCCACAGACTTGGACCAGCAATGCGACGGGCAGGATCTAGACCACGTCCCCAGTCTAGACTCGGCGGAGTCTGAGGAGGATCACGACGACCAAGACGATCATCTATTGTGA
>DPOGS214468-PA
MTDARTAQIPQRGSCKRKEDMASDTLPSPKKPRLVFTDLQRRTLQAIFKETKRPSKEMQVTIARQLGLEPTTVGNFFMNARRRSMDKWKDDDAPSTDLDQQCDGQDLDHVPSLDSAESEEDHDDQDDHLL-