| DPOGS214752 | ||
|---|---|---|
| Transcript | DPOGS214752-TA | 621 bp |
| Protein | DPOGS214752-PA | 206 aa |
| Genomic position | DPSCF300022 + 747023-767742 | |
| RNAseq coverage | 80x (Rank: top 64%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL021455 | 2e-81 | 90.51% | |
| Bombyx | BGIBMGA004747-TA | 2e-69 | 83.23% | |
| Drosophila | Nrx-1-PB | 5e-75 | 65.53% | |
| EBI UniRef50 | UniRef50_Q7PSM9 | 2e-78 | 77.65% | AGAP004066-PA n=4 Tax=Endopterygota RepID=Q7PSM9_ANOGA |
| NCBI RefSeq | XP_309618.4 | 1e-79 | 77.35% | AGAP004066-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|347971168 | 6e-78 | 77.65% | AGAP004066-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastx | gi|347971168 | 7e-77 | 70.19% | AGAP004066-PA [Anopheles gambiae str. PEST] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | nvi:100120851 | 5e-66 | ||
| K07377 (NRXN) | maps-> | Cell adhesion molecules (CAMs) | ||
| InterPro domain | [32-187] IPR008985 | 7.6e-43 | Concanavalin A-like lectin/glucanase | |
| [32-185] IPR013320 | 4.6e-36 | Concanavalin A-like lectin/glucanase, subgroup | ||
| [44-186] IPR001791 | 2.5e-34 | Laminin G domain | ||
| [52-184] IPR012680 | 8.7e-33 | Laminin G, subdomain 2 | ||
| Orthology group | MCL18586 | Insect specific | ||
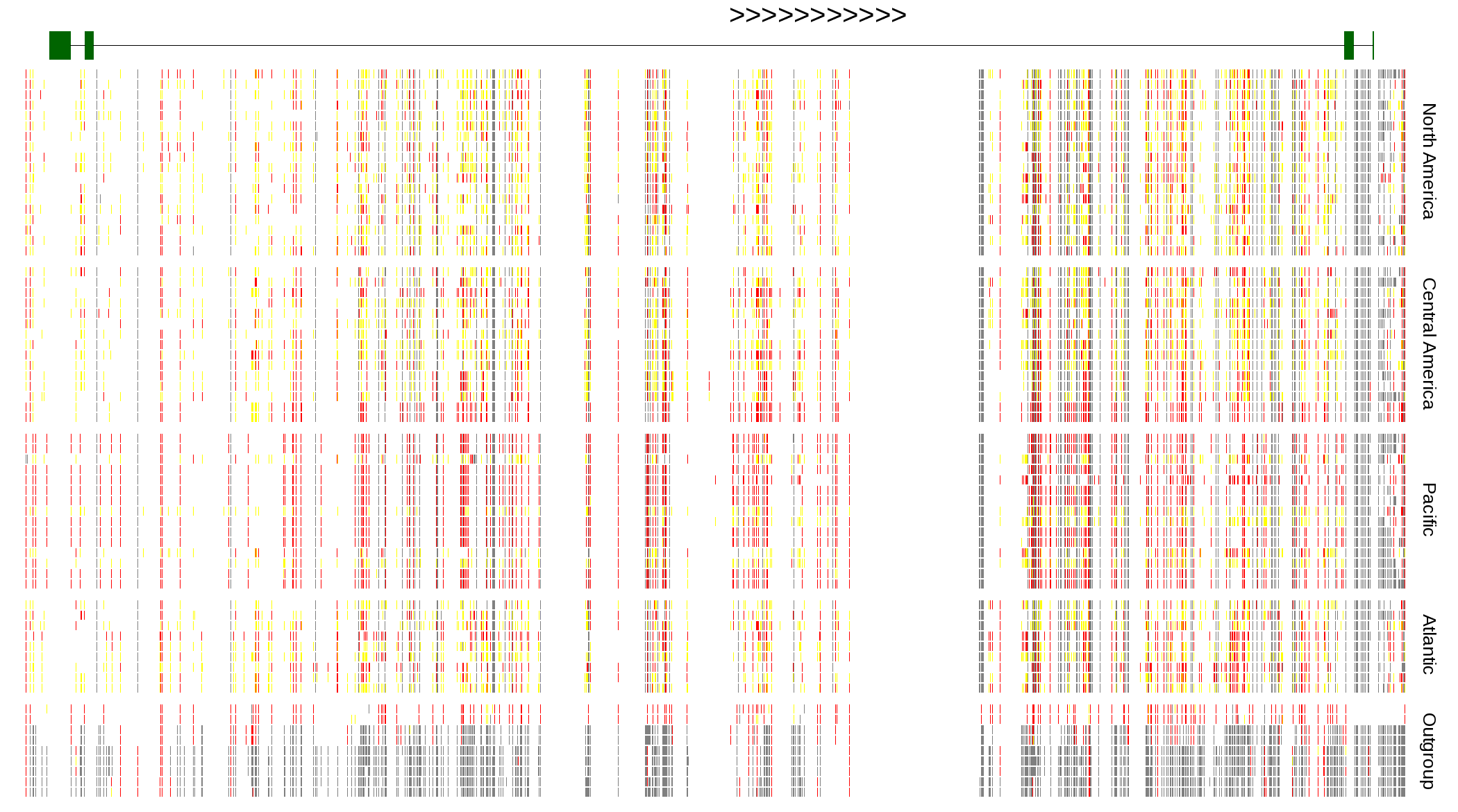
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214752-TA
ATGTCGGGACAGGTCAGAACGATGCGCTACATACTCTTCTGTCTTCTCTCCCTCTCGCTCAATAGGAACCTCGCGTTCGTCCTCGACAAACAGAACCCCTACTCCCAATTCCGCAAATGGAACGCCGGTTTGAACGGCACCTTGGAACTCGAGTTCAAAACCGATCAGCCGAACGGACTGCTATTGTACACGGACGACGGCGGCACTTACGACTTCTTCGAACTAAAGCTGGTGAATGGAGCCCTAAGACTGAGGTACAACTTGGGCGGCGGAGCTCAGATTATAACGGTGGGCAACAATTTAAACGACGGCCACTGGCACAAAGTTCAGGTGGGTCGTCGCGATGAACACACGTCTCTGTCGGTGGACGGCAGCACTCAGAGCAAGGCCTCCCGCGGCAAGGAGTTTGATTTCGGCAAGTTTAATACAAACTCGGATGTCTTCATCGGAGGGATACCGTCCTCGTACAATTCCAAACTGACGACCTTGGCTCTGCCGTCAGTGATTTTTGAGCCCAAGTTTCGTGGCTCAGTTCGCAACCTGGTGTACTCCGACTTACCGGGACAACCTCCTCGCAGACAAGAGCTACGACACTCTAGAGACTTGAAGTACGGGGTCTAA
>DPOGS214752-PA
MSGQVRTMRYILFCLLSLSLNRNLAFVLDKQNPYSQFRKWNAGLNGTLELEFKTDQPNGLLLYTDDGGTYDFFELKLVNGALRLRYNLGGGAQIITVGNNLNDGHWHKVQVGRRDEHTSLSVDGSTQSKASRGKEFDFGKFNTNSDVFIGGIPSSYNSKLTTLALPSVIFEPKFRGSVRNLVYSDLPGQPPRRQELRHSRDLKYGV-