| DPOGS215639 | ||
|---|---|---|
| Transcript | DPOGS215639-TA | 597 bp |
| Protein | DPOGS215639-PA | 198 aa |
| Genomic position | DPSCF300041 - 1794868-1795823 | |
| RNAseq coverage | 32x (Rank: top 75%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL014093 | 2e-87 | 75.52% | |
| Bombyx | BGIBMGA003538-TA | 1e-55 | 56.52% | |
| Drosophila | CG32196-PB | 1e-27 | 41.67% | |
| EBI UniRef50 | UniRef50_UPI0001A5CE84 | 2e-29 | 42.68% | UPI0001A5CE84 related cluster n=2 Tax=unknown RepID=UPI0001A5CE84 |
| NCBI RefSeq | NP_001155144.1 | 5e-30 | 42.68% | gamma-glutamyl cyclotransferase-like venom protein isoform 1 [Nasonia vitripennis] |
| NCBI nr blastp | gi|383865799 | 1e-29 | 46.05% | PREDICTED: gamma-glutamylcyclotransferase-like [Megachile rotundata] |
| NCBI nr blastx | gi|239735532 | 3e-29 | 42.68% | gamma-glutamyl cyclotransferase-like venom protein isoform 2 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003839 | 2.2e-37 | gamma-glutamylcyclotransferase activity | |
| KEGG pathway | cfa:611603 | 8e-22 | ||
| K00682 (GGCT) | maps-> | Glutathione metabolism | ||
| InterPro domain | [5-150] IPR017939 | 2.2e-37 | Gamma-glutamylcyclotransferase | |
| [3-154] IPR013024 | 7.5e-36 | Butirosin biosynthesis, BtrG-like | ||
| Orthology group | MCL30178 | Lepidoptera specific | ||
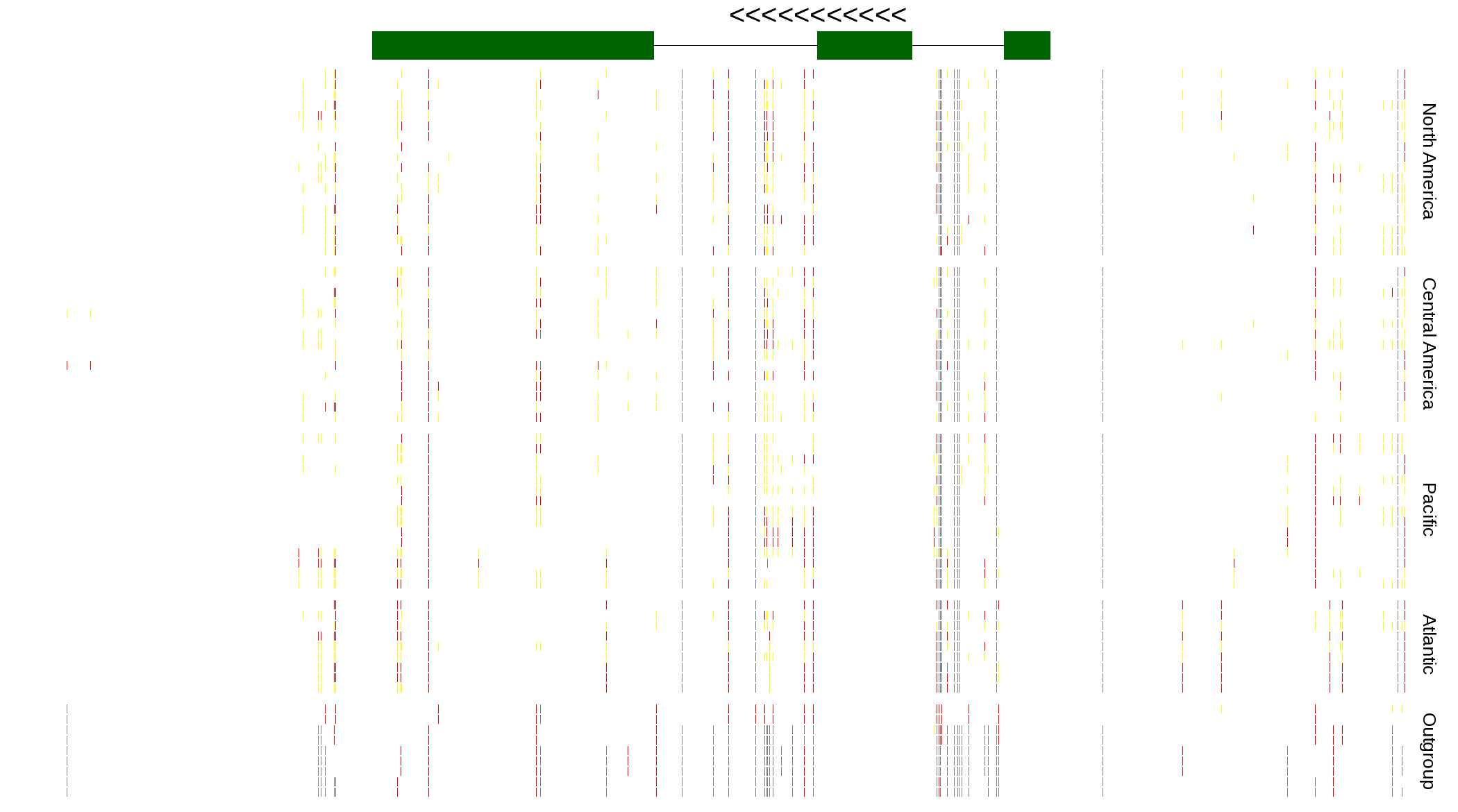
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS215639-TA
ATGTTTTCGTTTAGAATACACATGAATAATCCATCAGCGGAATTTGTATCAATAGCAAGACTTGATAATTACAGGTTAGATTTCATAAAGTATTCCAAATTCTGGGGAGGACCTACGGCTACCATCGTGCCGACTGCGAACGCCCAAGTGTGGGGCGTTATATGGCGCTTGGACGTAGACAAATTGTCGATTTTGGATGAGCAAGAGGGCGTAGAACGTAAAATTTACTACCCAACCCACGTTCAAGTACTCACACCTTACATGGGGTCTTTTACTTGCCGTGTTTACATCCACAAGGTCAATCCGTTGCCTAGAGGAGACAATGACGTCATACCTGTCGAGCGCTGGCCTTCTTGGACCTACAAGGAAGTTATCATTATAGGTGCAATGGAGCACGAGCTGCCAGAATACTATATACAGAGTTTAGAAAAAATTAAAGACAACGGGGAAAGGGGATGCCTCAAGATGTATAGTCTACTGATACGATATGCTAATGACGCTCCTTGCGTCTGTCCTCCCCCGCGCCGGATACCGAAACCACCAGTATTAGACTTGAGAGCATTGAGGGAGCGGAACAGAGAGGGGGTTGAAGATTAA
>DPOGS215639-PA
MFSFRIHMNNPSAEFVSIARLDNYRLDFIKYSKFWGGPTATIVPTANAQVWGVIWRLDVDKLSILDEQEGVERKIYYPTHVQVLTPYMGSFTCRVYIHKVNPLPRGDNDVIPVERWPSWTYKEVIIIGAMEHELPEYYIQSLEKIKDNGERGCLKMYSLLIRYANDAPCVCPPPRRIPKPPVLDLRALRERNREGVED-