| DPOGS216029 | ||
|---|---|---|
| Transcript | DPOGS216029-TA | 327 bp |
| Protein | DPOGS216029-PA | 108 aa |
| Genomic position | DPSCF300067 - 621200-622295 | |
| RNAseq coverage | 4966x (Rank: top 3%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL008949 | 5e-58 | 96.30% | |
| Bombyx | BGIBMGA009012-TA | 1e-57 | 95.37% | |
| Drosophila | Cyt-c-p-PA | 5e-52 | 85.19% | |
| EBI UniRef50 | UniRef50_G7Y2H8 | 8e-46 | 76.64% | Cytochrome c n=4 Tax=Opisthokonta RepID=G7Y2H8_CLOSI |
| NCBI RefSeq | XP_310154.1 | 5e-51 | 86.11% | AGAP009537-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|124076981 | 1e-54 | 95.37% | RecName: Full=Cytochrome c |
| NCBI nr blastx | gi|124076981 | 1e-55 | 95.37% | RecName: Full=Cytochrome c |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0009055 | 2.2e-81 | electron carrier activity | |
| GO:0020037 | 2.2e-81 | heme binding | ||
| GO:0005506 | 2.2e-81 | iron ion binding | ||
| KEGG pathway | aga:AgaP_AGAP009537 | 2e-50 | ||
| K08738 (CYC) | maps-> | Colorectal cancer | ||
| Amyotrophic lateral sclerosis (ALS) | ||||
| Viral myocarditis | ||||
| Alzheimer's disease | ||||
| Apoptosis | ||||
| Small cell lung cancer | ||||
| Huntington's disease | ||||
| Pathways in cancer | ||||
| p53 signaling pathway | ||||
| Parkinson's disease | ||||
| InterPro domain | [1-107] IPR002327 | 2.2e-81 | Cytochrome c, class IA/ IB | |
| [5-107] IPR009056 | 1.2e-45 | Cytochrome c domain | ||
| [8-105] IPR003088 | 1.1e-11 | Cytochrome c, class I | ||
| Orthology group | MCL11053 | Single-copy universal gene | ||
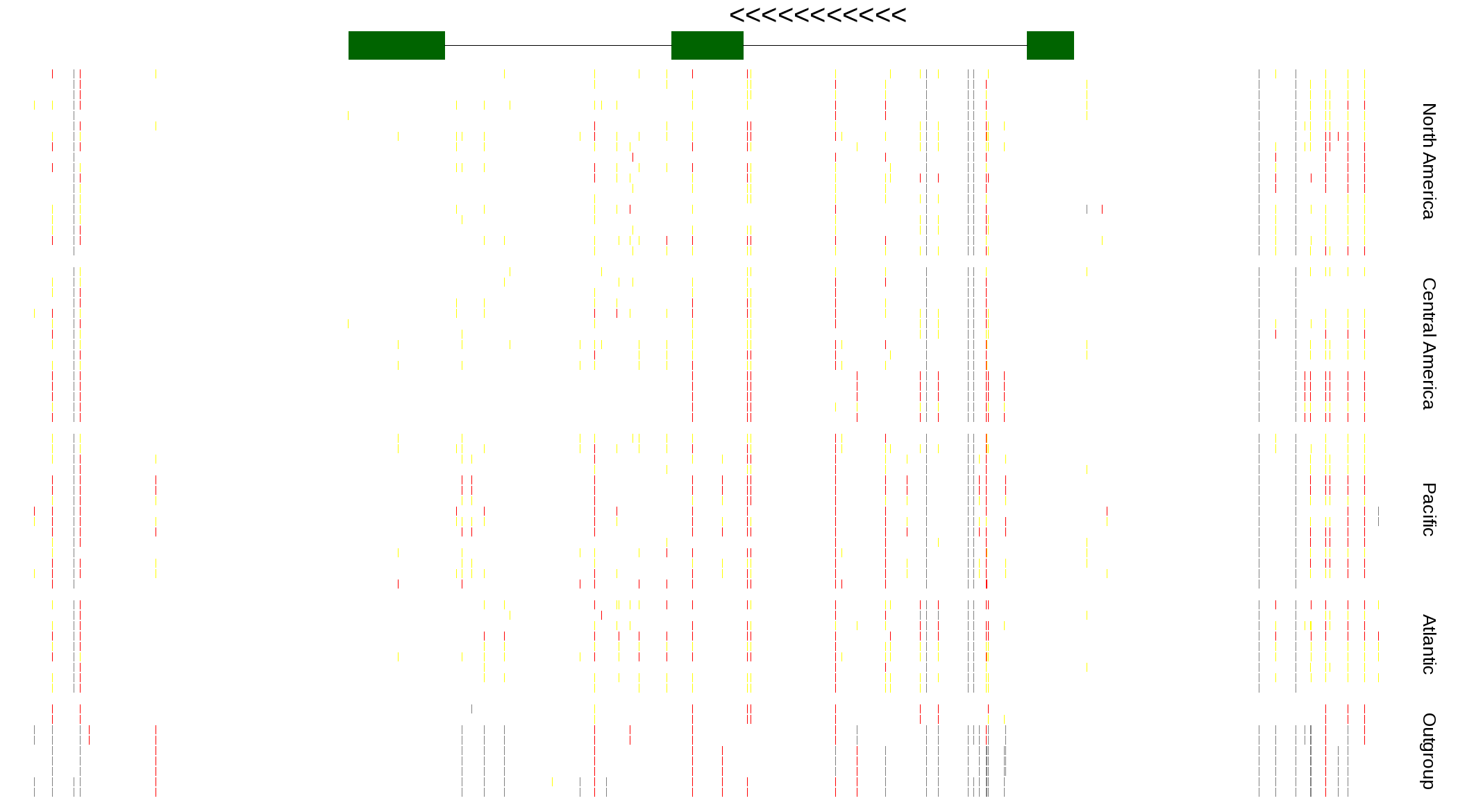
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS216029-TA
ATGGGTGTCCCTGCTGGAAATGCGGAAAACGGCAAAAAGATCTTTGTTCAAAGATGTGCTCAATGTCACACCGTTGAGGCTGGCGGCAAGCACAAAGTAGGACCCAACCTACATGGTTTCTTTGGTCGCAAAACTGGTCAAGCTTCCGGGTTTAACTATTCAGAAGCCAACAAAGCTAAAGGTATCACCTGGGGCGATGACACACTCTTTGAGTACTTGGAGAACCCTAAGAAATACATCCCCGGAACAAAGATGGTGTTTGCTGGCCTCAAGAAAGCAAATGAACGTGCGGATCTAATTGCCTACCTTAAAGAAGCTACTAAGTAA
>DPOGS216029-PA
MGVPAGNAENGKKIFVQRCAQCHTVEAGGKHKVGPNLHGFFGRKTGQASGFNYSEANKAKGITWGDDTLFEYLENPKKYIPGTKMVFAGLKKANERADLIAYLKEATK-