| DPOGS216208 | ||
|---|---|---|
| Transcript | DPOGS216208-TA | 597 bp |
| Protein | DPOGS216208-PA | 198 aa |
| Genomic position | DPSCF300080 + 517744-518340 | |
| RNAseq coverage | 102x (Rank: top 61%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL015841 | 2e-57 | 61.11% | |
| Bombyx | BGIBMGA004515-TA | 4e-69 | 64.09% | |
| Drosophila | l(2)efl-PA | 5e-40 | 48.59% | |
| EBI UniRef50 | UniRef50_Q5R1P4 | 3e-67 | 64.09% | Heat shock protein hsp23.7 n=17 Tax=Pancrustacea RepID=Q5R1P4_BOMMO |
| NCBI RefSeq | NP_001036942.1 | 6e-68 | 64.09% | heat shock protein hsp23.7 [Bombyx mori] |
| NCBI nr blastp | gi|112983144 | 9e-67 | 64.09% | heat shock protein hsp23.7 precursor [Bombyx mori] |
| NCBI nr blastx | gi|112983144 | 6e-64 | 61.03% | heat shock protein hsp23.7 precursor [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | dme:Dmel_CG4533 | 3e-38 | ||
| K09542 (CRYAB) | maps-> | Protein processing in endoplasmic reticulum | ||
| InterPro domain | [76-170] IPR002068 | 2.1e-26 | Heat shock protein Hsp20 | |
| [27-39] IPR001436 | 2.6e-25 | Alpha crystallin/Heat shock protein | ||
| [62-171] IPR008978 | 2.2e-16 | HSP20-like chaperone | ||
| Orthology group | MCL30260 | Lepidoptera specific | ||
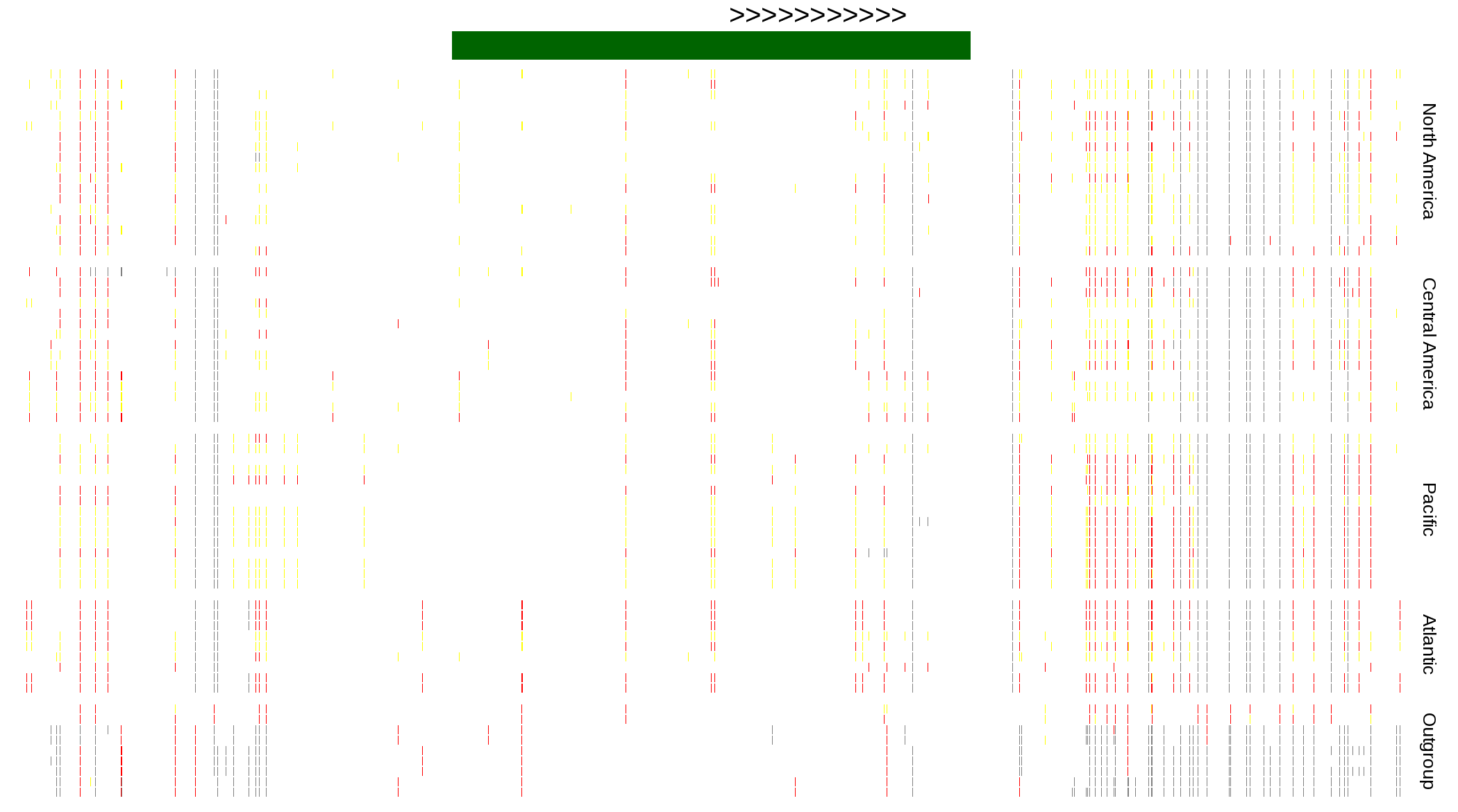
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS216208-TA
ATGGATAAGCTAGTTTATTTACTTTTCATCACTGGTATTCTAACTGGTGCATACTGTGACGAAGAATGTTCAAGAAAGCGAAGGAACAGTTTACTCGATCAAGATTTCGGTATGTCATTGACGGATGACGATTTAATCACCACAATGATGGCTCCTTTGATGTTAAGAAACTACTTCAGACCCTGGCGCTACCTTGAACCAATGGCAAGGGATATCGGTTCCACTATCAAGACGGATAAGGATAAATTTACGATCAACGTCGATGTTCAACATTTCGCACCTGACGAAATAACTGTAAGAACTGCGGAGGGTTACGTTATAGTTGAAGCGAAACATGAAGAGAAGCAAGACGAACACGGCTTTGTTTCTCGTCAGTTCATGAGGAGATACTCATTGCCAGAAGGTGTCGAGTCTGAAGATGTTTTATCAGAGCTCTCGAGCGACGGCATCTTGACAATCTCGGCTCCAAGGAAAGATGTTGATAAAAAAGGAGAGAGAATTGTTACAATAACTAAGACTGGACCCGTGCGTAAACAAGCAAAGGAGACGAAAGAGGAAAGTTGCTCTGCCGATACTTGTGAAAAACCTAAAGAATAA
>DPOGS216208-PA
MDKLVYLLFITGILTGAYCDEECSRKRRNSLLDQDFGMSLTDDDLITTMMAPLMLRNYFRPWRYLEPMARDIGSTIKTDKDKFTINVDVQHFAPDEITVRTAEGYVIVEAKHEEKQDEHGFVSRQFMRRYSLPEGVESEDVLSELSSDGILTISAPRKDVDKKGERIVTITKTGPVRKQAKETKEESCSADTCEKPKE-